137 human genomes from the Middle East fill gaps in human history

Whole-genome sequencing efforts around the world have offered important insights into human diversity, historical migrations, and the relationships between people of different regions—but scientists still don't have a complete picture because some regions and people remain understudied. A new study reported in the journal Cell on August 4 helps to fill one of these big gaps by generating more than 100 high-coverage genome sequences from eight Middle Eastern populations using linked-read sequencing.
"The Middle East is an important region to understand human history, migrations, and evolution: it is where modern humans first expanded out of Africa, where hunter-gatherers first settled and transitioned into farmers, where the first writing systems developed, and where the first major known civilizations emerged," says Mohamed Almarri of the Wellcome Sanger Institute, UK. "However, despite this importance, the region has been historically understudied in genomic studies."
In the new study, Almarri, Marc Haber (University of Birmingham, UK), and their colleagues sequenced 137 whole genomes from eight Middle Eastern populations.
By generating the most comprehensive resource of human genetic variation in the Middle East using a new sequencing technology called linked-read sequencing, the researchers were able to reconstruct the genomic history of the region with unprecedented resolution. The researchers say that some of the events recorded in the Middle Eastern genomes could be linked with what's known from archeology or linguistics, such as the invention of agriculture and the spread of Semitic languages. But other events can only be elucidated by studying the DNA of ancient and modern people who lived in the region.
Some of their most notable findings include the following:
- The identification of 4.8 million new gene variants that are specific to Middle Eastern populations that could now provide the basis for future research.
- Genetic variants that show evidence of selection—in other words, mutations that spread unusually quickly—potentially due to adaptation to the changing environment and lifestyle.
- In the Levant, where agriculture was first developed, populations experienced a massive growth around the transition to agriculture that wasn't paralleled in Arabia.
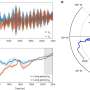
- Arabian populations suffered a severe population decrease around 6,000 years ago, which coincides with the change in climate in Arabia turning it from a green, wet region into the largest sand desert in the world today.
- Middle Easterners descend from the same population that expanded out of Africa 50,000 to 60,000 years ago.
- Arabian groups have significantly lower Neanderthal ancestry than other Eurasians, potentially caused by excess basal Eurasian and African ancestry in Arabians that depletes their Neanderthal ancestry
- The movement of populations during the Bronze Age potentially spread the Semitic languages from the Levant to Arabia and East Africa.
- An increase in the frequency of variants associated with type 2 diabetes in some populations in the past 2,000 years, suggesting that variants that were beneficial in the past are today associated with diseases.
"We found 4.8 million variants that were not previously discovered in other populations," Haber says. "Hundreds of thousands of these are common in the region, and any of them could hold medical relevance."
"Our study fills a major gap in international genomic projects by cataloguing genetic variation in the Middle East," says Chris Tyler-Smith of the Wellcome Sanger Institute, UK. "The millions of new variants we found in our study will improve future medical association studies in the region. Our results explain how the genetics of Middle Easterners formed over time, providing new insights, which complement knowledge from archeology, anthropology, and linguistics."
The researchers say they will now follow up on variants that show evidence of selection. Through these continued studies, they hope to further understand the biological effects of those newly found variants while further refining the genetic history of the region.
More information: Cell, Almarri et al.: "The Genomic History of the Middle East" www.cell.com/cell/fulltext/S0092-8674(21)00839-4 , DOI: 10.1016/j.cell.2021.07.013
Journal information: Cell
Provided by Cell Press