First-ever gene expression map of an entire nervous system completed
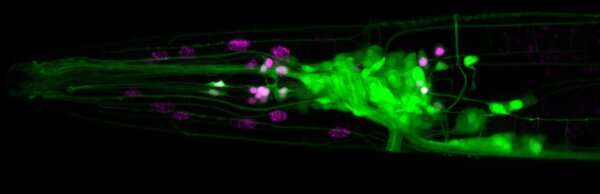
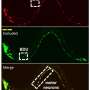
Research Assistant Professor Seth Taylor and Professor David Miller, both in Vanderbilt University's Department of Cell and Developmental Biology, have established a gene expression atlas for the nervous system of the nematode C. elegans, along with biologists from Colombia University and Yale University.
Their data complement the known wiring diagram of the C. elegans nervous system to create, for the first time, a complete picture of gene expression for every neuron in an entire nervous system.
The researchers used flow cytometry methods developed at Vanderbilt to capture every type of neuron for gene expression profiling and have shared these data for anyone to investigate how individual genes in neurons contribute to the function of a nervous system. To illustrate this approach, they used computational methods to identify DNA regions that control gene expression in specific neurons, as well as adhesion proteins that may sustain the formation of synapses—the connections between neurons from which all brain activity initiates.
While the human brain has 100 billion neurons, C. elegans has only 302. "Because the genetic rules that direct the development and function of the worm nervous system are also likely to operate in the brain, we expect that this unique gene expression data set will serve as a valuable foundation for deciphering the genetic underpinning of the brain," Miller said. "These data will facilitate hypothesis-driven research into how genes specify different neuron types, how they make and maintain connections, and how they influence behavior."
By sharing their data, Miller and colleagues are aiming to create opportunities for researchers studying the nervous system and how genetic defects shape the brain's function and design. "We expect that our findings will spur these long-term efforts," Miller said.
The article "Molecular topography of an entire nervous system" was published online in the journal Cell on July 7.
More information: Seth R. Taylor et al, Molecular topography of an entire nervous system, Cell (2021). DOI: 10.1016/j.cell.2021.06.023
Journal information: Cell
Provided by Vanderbilt University