Starving tuberculosis of sugars may be a new way to fight it
Tuberculosis is a devastating disease that claims over 1.5 million lives each year. The increase in TB cases that are resistant to the current antibiotics means that novel drugs to kill Mycobacterium tuberculosis (Mtb) are urgently needed. Researchers from the University of Warwick have successfully discovered how Mycobacterium tuberculosis uses an essential sugar called trehalose, which provides a platform to design new and improved TB drugs and diagnostic agents.
Tuberculosis (TB), caused by the bacterial pathogen Mycobacterium tuberculosis (Mtb) is the leading cause of death from a single infectious agent world-wide claiming over 1.5 million lives each year.
Mycobacterium tuberculosis (Mtb) is a very unique pathogen and is able to survive in the human body for decades. One way that Mtb survives is by 'eating' scarce energy sources for nutrition, whilst at the same time the human host attempts to limit the food that is available.
However, we need a better understanding of Mtb's intracellular diet because inhibiting the pathways that allow Mtb to access and use essential food sources could be good targets for the development of new anti-tubercular agents.
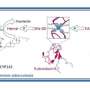
One source of energy that Mtb uses is a sugar that is found in its own cell wall, called trehalose. It appears that Mtb has evolved a unique strategy to recycle and reuse this sugar to ensure that it does not waste any potential energy sources, which are in short supply.
The transport protein, which is responsible for the uptake of trehalose, called LpqY, is essential for Mtb to establish infection. If the LpqY protein is deleted and no longer able to function then Mtb can no longer supply itself with trehalose and becomes less pathogenic.
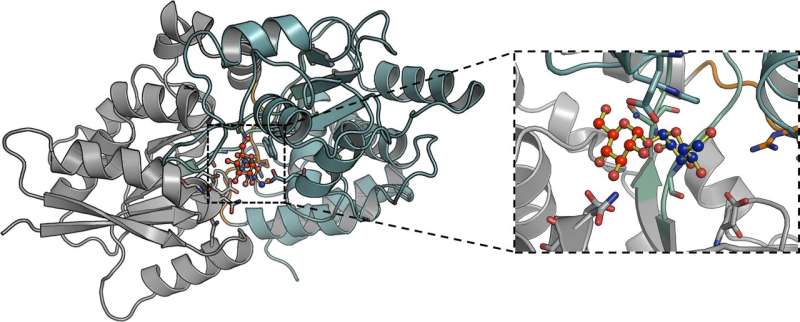
In the paper, "Structural basis of trehalose recognition by the mycobacterial LpqY-SugABC transporter," published in the Journal of Biological Chemistry, researchers from the School of Life Sciences at the University of Warwick, have unraveled the molecular basis of how Mtb uses and transports trehalose, a process which is specific to Mtb and does not occur in humans.
The team used X-ray crystallography to determine the 3-dimensional structure of LpqY and analyzed how this important transport protein is able to bind and recognize trehalose. They then went on to use a number of experimental techniques which showed that LpqY is highly specific for trehalose, is also able to recognize sugars that are similar to trehalose with small modifications and map key recognition features.
Dr. Elizabeth Fullam, who is a Sir Henry Dale Fellow from the School of Life Sciences at the University of Warwick, said, "It is vital that we find new innovative strategies to combat TB. The LpqY trehalose transporter is a potential drug target because when it is not functioning it results in Mtb becoming less virulent. Now that we understand exactly how trehalose is recognized we will be able to design specific molecules that will enable us to kill TB. Alternatively, another possibility is that we can use the LpqY transporter to our advantage and find ways to deliver compounds for TB diagnosis."
More information: Christopher M. Furze et al. Structural basis of trehalose recognition by the mycobacterial LpqY-SugABC transporter, Journal of Biological Chemistry (2021). DOI: 10.1016/j.jbc.2021.100307
Journal information: Journal of Biological Chemistry
Provided by University of Warwick