Study proves bits of DNA in seawater correlate to the weight of netted fish
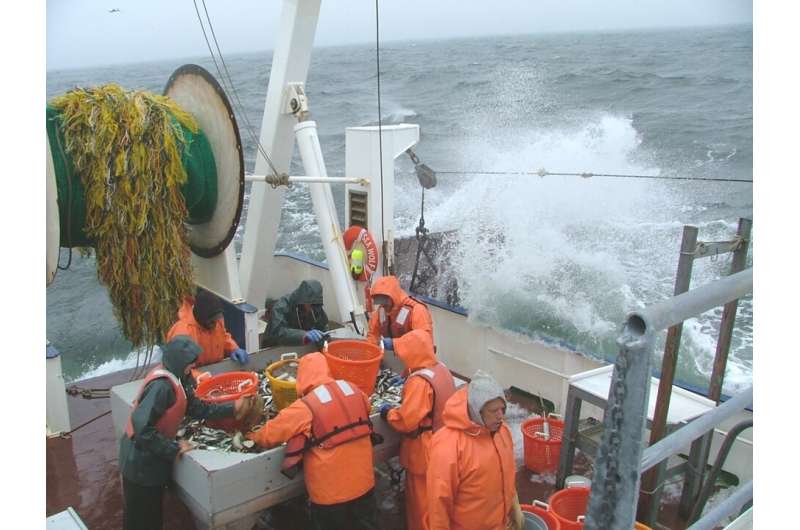

Humanity is a step closer to answering one of the most ancient of questions—"how many fish in the sea?"—thanks to newly-published proof that the amount of fish DNA collected in a water sample closely corresponds to kilos of fish captured in a trawl with nets.
In a breakthrough study, scientists report that floating bits of DNA found in small water samples reveal the relative biomass of fish in the sea roughly as well as a 'gold standard' US state government trawl with nets.
The researchers drew seawater samples during New Jersey government fish trawls and tested the water for fish DNA. Analysis of the water was able to reveal the relative abundance of fish with a 70% match in results between the two sampling methods. In addition to the great concordance between methods, the study found that each sampling method yielded information missed by the other.
While environmental DNA ("eDNA") has been proven before as a reliable way to determine the variety of fish in an area of water, the new study is the first to show that bits of eDNA floating in seawater also disclose the relative abundance of the species swimming through it.
Published by the prestigious ICES Journal of Marine Science, the paper certifies 'fishing for DNA' as an inexpensive, harmless complement to nets, acoustics and other established ways to monitor the health of fish stocks and/or the shifting diversity, distribution and abundance of aquatic life.
The paper, a collaboration between The Rockefeller University, Monmouth University, and the New Jersey Bureau of Marine Fisheries, says the information about the diversity and relative abundance of fish available in a one-liter sample is comparable to a 66 million liter trawl sweep, enough seawater to fill a football stadium to the top of the goalposts.
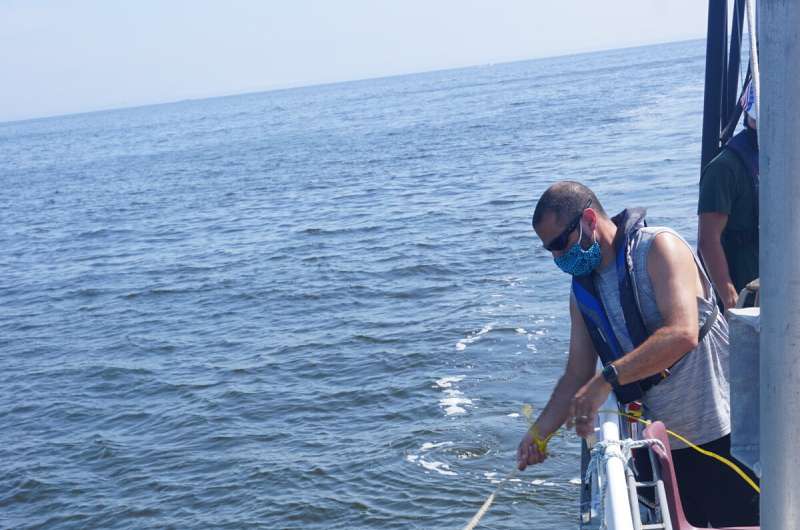
During four voyages by the New Jersey Ocean Trawl Survey in 2019 aboard the research vessel Sea Wolf, scientists led by Dr. Mark Stoeckle, Senior Research Associate at The Rockefeller University Program for the Human Environment, drew one-liter pop-bottle sized water samples from various depths just before the trawler's nets were lowered.
The finding has profound implications for improving global fisheries management and has led to early proposals for a "Great American Fish Count" in rivers and coastal waters, aided by millions of citizen scientists, comparable to Audubon's Great Backyard Bird Counts.
Fish and other organisms shed DNA like dandruff, Dr. Stoeckle explains, leaving an invisible trail wherever they swim. This environmental DNA can be skin cells, droppings, urine, eggs, and other biological residues that last in the ocean for a few days.
One year of eDNA sampling, out-of-pocket costs: $12,000
The eDNA process is straightforward and extremely inexpensive compared with traditional marine life monitoring methods, which involve ships with large crews and hand counts.
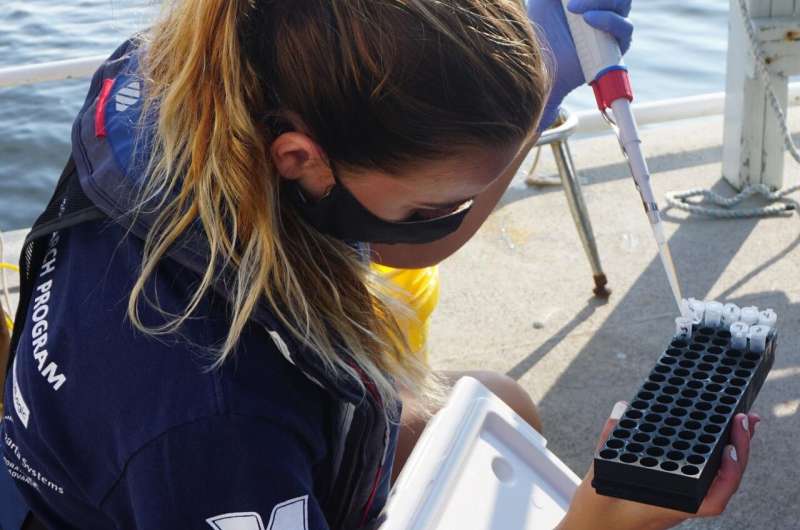
Co-author Zachary Charlop-Powers at The Rockefeller University, lead developer of the software used in the DNA analyses, explains that eDNA testing involves collecting and filtering a water sample, extracting and sequencing the DNA in a laboratory, then matching the results found in an online DNA reference library.
"The bioinformatic tools used by the team are the same 'barcode' analysis pipelines commonly used by microbiologists but were adapted for the study of marine vertebrates."
He notes too that the year of sampling and DNA extraction required an investment of just $12,000, exclusive of salaries.

"The applications of environmental DNA in the marine realm are vast," says Dr. Stoeckle, a Harvard-educated MD who helped pioneer DNA "barcoding," the identification of species from a small region of the animal's DNA sequence.
"eDNA offers a low-cost way to monitor the effectiveness of a marine protected area, for example, or whether efforts to restore a coral reef are succeeding. It could reveal the ecological effects of marine industrial activities, including offshore wind farms, oil and gas rigs, and commercial and recreational fishing."
Adds Dr. Stoeckle: To put this in perspective, if we thought of a trawl as a full medical CAT or MRI scan, then eDNA can be thought of as a pocket ultrasound—it can be carried and used anywhere in the hospital, without the time and expense of scheduling a full-scale exam. And eDNA surveys will become better and more informative every year as the technique improves and the DNA reference library grows.
Says co-author Dr. Jason Adolf, Endowed Associate Professor of Marine Science, Monmouth University, "eDNA could also be used to identify life in ocean regions hard to access with trawls, such as very rocky areas, or places too deep or too shallow."
Monmouth co-author Dr. Keith J. Dunton, an expert on endangered fish species, notes that the results are promising for rare as well as common fish species.
"eDNA along with other technologies like acoustic telemetry offers a sensitive, non-extractive way to monitor declines and revivals of rare, threatened, and endangered species," he says. "We do not have to put them through stressful capturing to know that they are there."
Trawl surveys, the main tool used to monitor fish populations, have carefully established protocols and yield rich information but are costly, time-consuming, and require special equipment and fish identification experts. Due to the crew size needed, such trawls have been limited recently by COVID-19.
The New Jersey surveys every season involve deploying a bottom trawl, similar to that used in commercial fishing, behind a vessel over a predetermined pattern. The catches in the nets are hauled up and sorted on tables where the weight of each identified species is recorded. Between 30 and 40 trawls are done about every three months.
To compare the trawl survey to the eDNA survey, one-liter water samples were collected at the surface and at depth before the trawls were done. However, samples were only taken before every fourth trawl. When the data from the two surveys were analyzed, the eDNA survey found most of the same fish species, and also found species not captured in the trawl. And it did so with only one-quarter of the samples taken and a fraction of the effort involved.
The paper says most (70% to 87%) species detected by trawl in a given month were also detected by eDNA, and vice versa, including nearly all (92% to 100%) abundant species. Conversely, most dropouts were relatively rare taxa.
Trawl and eDNA peak seasonal abundance agreed for about 70% of fish species.
In other comparisons, monthly eDNA species "reads" correlated with the monthly weight, or biomass, of that species recovered in the trawl.
The eDNA reporting "largely concorded with monthly trawl estimates of marine fish species richness, composition, seasonality, and relative abundance," the paper says.
"It's important to understand that the results of both methods are true, and complementary," noted Stoeckle. "They catch a lot of overlapping, concordant information as well as some information unique to each method."

Gregory Hinks of the New Jersey Department of Environmental Protection, who co-authored the paper with Bureau of Marine Fisheries colleague Stacy M. VanMorter, adds: "During times like COVID when it is unsafe to conduct surveys with large crews, the eDNA method might allow us still to maintain some continuity in our surveys. In any case, piggybacking eDNA onto an existing survey may eventually provide an affordable way to improve marine fish stock assessment."
The new paper lays out further research required, such as better calibration of eDNA 'reads' to fish body mass—how much DNA is shed by 1,000 anchovies weighing 1 kilo, for example, compared with a one kilo sized sea bass?—and how to account for eDNA reads that may be the result of injury due to a predator attack.
Since collecting water for eDNA is so quick and easy to do, research or oceanographic vessels and commercial and recreational vessels can collect samples as they travel from place to place. Even drones could be deployed to collect water samples.
And with the benefit of additional studies in marine and freshwaters, estimates of animal numbers using eDNA will continue to improve as well as the DNA reference data banks that allow reliable identification of aquatic species.
eDNA opens the way to surveys of unprecedented value, quality, and affordability, says Jesse Ausubel, Director of The Rockefeller University's Program for the Human Environment, who developed and helped oversee the first international Census of Marine Life, a decadal (2000-2010) collaboration of about 2,700 scientists in 80 countries.
"eDNA makes the ocean a sea of biological information," he says. "In the U.S. we could organize a Great American Fish Count in which millions of citizen scientists might collect water for eDNA testing spanning all our waters. Globally, the incipient UN Decade of the Oceans could include a Great Global Fish Count sampling from sea floor to sea surface and near shore to mid-ocean all during a single day or week."
Tony MacDonald, Director of the Monmouth University Urban Coast Institute, says "Our institute and scientists were excited to support this innovative work, one of several partnerships in recent years between UCI and The Rockefeller University Program for the Human Environment."
"We hope to have the opportunity to continue and expand our collaboration with New Jersey's Department of Environmental Protection Marine Fisheries and the National Oceanic and Atmospheric Administration on future fish trawls to further advance eDNA research."
Comments Tim Gallaudet, Ph.D., Rear Admiral, U.S. Navy (Ret.) Assistant Secretary of Commerce for Oceans and Atmosphere and Deputy NOAA Administrator: "NOAA is rapidly advancing 'omics technologies, including eDNA, to improve our ability to monitor and understand biological communities in our oceans and the Great Lakes."
"Important applications include monitoring endangered and invasive species, assessing biodiversity for ecosystem health, tracking aquaculture pathogens, and augmenting fisheries surveys."
"Through the NOAA 'Omics Strategy' and our forthcoming Implementation Plan, we have defined goals and actionable steps to integrate modern 'omics technologies to help meet our mission. Collaboration with Rockefeller University and other partners will allow us to expand and advance 'omics research and eDNA in direct support of the American Blue Economy."
Marine eDNA's potential applications include:
- Exploration: discovering species previously unknown in certain ranges
- Discovering rare species and others unknown to science (or absent from genome databases)
- Sampling remote, difficult-to-reach, and intriguing places
- Assessment of the size of fish stocks
- Identifying the range of marine animals
- Determining the effect of protected area designation on fish and other marine animal populations and other forms of ecological restoration
- Monitoring the effect on native species of fish farming operations, offshore oil and gas operations, or wind farms
- Determining the effects of artificial reefs, of severe storms and other disturbances to marine ecosystems including harmful algal blooms
- Monitoring vulnerable, threatened or endangered species, invasive species, or the presence of species dangerous to swimmers
- Gauging the impacts of climate variability
- Mapping marine animal diversity, distribution, migration and abundance, including invasive species, and species popular with sport fishers
More information: Mark Y Stoeckle et al. Trawl and eDNA assessment of marine fish diversity, seasonality, and relative abundance in coastal New Jersey, U.S.. ICES Journal of Marine Science.Published: 3 December 2020. doi.org/10.1093/icesjms/fsaa225
Provided by ICES Journal of Marine Science