Researchers reveal structural basis for two metal-ion catalysis of DNA cleavage

Cas9 and Cas12a, two Cas proteins used most in gene-editing, both encompass a RuvC catalytic domain. To understand how the RuvC domain cleaves DNA, it is critical to elucidate the structures of RuvC-containing Cas complexes in their catalytically competent states, with both metal-ions and ssDNA substrate bound in the RuvC catalytic pocket.
Cas12i2, a Class 2 type V-I CRISPR-Cas, was discovered in 2018. With molecular weight lighter than Cas12a and Cas9, Cas12i2 is expected to be an emerging tool for gene editing. To learn more about its structure and catalytic mechanism is of great significance.
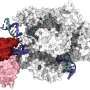
In a study published in Nature Communications, the researchers from the Institute of Biophysics of Chinese Academy of Sciences (CAS) and the University of Science and Technology of China of CAS revealed the crystal structures of Cas12i2-crRNA complexes and Cas12i2-crRNA-DNA complex, the mechanism of DNA recognition and cleavage by Cas12i2, and the activation of the RuvC catalytic pocket induced by a conformational change of the Helical-II domain.
The researchers reported high-resolution structures of Cas12i2-crRNA in three states, including the binding state, the seed region pairing state and the catalytic state. These data suggested that the crRNA:DNA duplex of 13-nt and longer has the ability to initiate cleavage, Helical-II domain conformational change activates the RuvC domain.
They then captured the catalytic state of Cas12i2, with both metal ions and the ssDNA substrate bound in the RuvC catalytic pocket, which revealed the essential roles of metal-ions for DNA cleavage by the RuvC domain in Cas12 and Cas9. This is also the first time an ssDNA and two metal ions were observed in the RuvC catalytic pocket, showing their active state.
Additionally, the structural and biochemical characterization of Cas12i2 provided insights into the molecular mechanisms of this CRISPR effector and suggested potential applications of Cas12i2. The researchers observed that pre-crRNA processing has effects on dsDNA cleavage activity. Biochemical studies showed that Cas12i2 cleaves ssDNA in a PAM-independent manner. Furthermore, the formation of the crRNA:target DNA hybrid activates the RuvC catalytic pocket, with base-pairing in the seed region not being essential for activation.
This study provided significant insights into the DNA cleavage mechanism by RuvC-containing Cas proteins, and had implications for developing more reliable genome-editing or nucleic acids detection tools.
Clustered regularly interspaced short palindromic repeats (CRISPR)-Cas (CRISPR-associated proteins) is an RNA-guided adaptive immunity system present in bacteria and archaea.
More information: Xue Huang et al. Structural basis for two metal-ion catalysis of DNA cleavage by Cas12i2, Nature Communications (2020). DOI: 10.1038/s41467-020-19072-6
Journal information: Nature Communications
Provided by Chinese Academy of Sciences