Research opens the way to new drugs

Research by a team at Te Herenga Waka–Victoria University of Wellington's School of Biological Sciences dispels the belief that on the assembly line of enzymes there is a "proof-reading" mechanism that ensures molecules are put together in a certain way.
This could pave the way to designing improved anti-cancer compounds or drugs that can treat what are currently antibiotic-resistant superbugs.
"This is quite a big deal, as it opens the way to rationally redesigning antibiotics at a genetic level to counter the spread of antibiotic-resistant superbugs, as well as making analogs of anti-cancer drugs to improve activity or reduce toxic side-effects," says Professor David Ackerley, the university's biotechnology program director.
"If we can efficiently make analogs of these compounds by re-engineering the genetic blueprints for the assembly line that makes them, then we can much more rapidly explore new and improved drug candidates. It also offers prospects for a much more cost-effective way of making any improved drugs we discover."
In their paper just published in Nature Communications, Professor David Ackerley and his colleagues Dr. Mark Calcott and Dr. Jeremy Owen show they can make new molecules without having to alter the part of the "assembly line that houses the hypothetical proofreading machinery."
They focus on a key mechanism by which many antibiotics are made.
"Most antibiotics currently used in the clinic are based on molecules produced by bacteria to kill other bacteria, to help them compete successfully for limited resources," Professor Ackerley says.
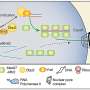
"Throughout evolution this has had a bit of an 'arms race' quality to it—bacteria make a new antibiotic to kill competing bacteria, the competing bacteria then develop resistance to the antibiotic, the first bacteria then modify their antibiotic so it can get around the resistance mechanisms, and so on. One of the aspects that helps bacteria quickly modify their antibiotics is that they build them using little nano-machines called enzymes—in our case 'non-ribosomal peptide synthetase' or 'NRPS' enzymes—which form an assembly line, with each enzyme adding a specific part of the final antibiotic molecule. To change part of the molecule, the producing bacteria don't need to evolve a whole new assembly line, but instead can just swap out the genes that encode a section of the assembly line for compatible genes that encode subtly different enzymatic machinery. This new machinery will then incorporate a different molecular part into the final antibiotic."
Professor Ackerley says for more than 20 years there has been a dominant belief that if scientists want to change part of the assembly line, they also need to swap a neighboring section believed to "proofread" the growing molecule.
"We performed a comprehensive analysis of how NRPS gene clusters have evolved in nature and found no evidence that such a proofreading mechanism exists. We also tried to generate new molecules by mimicking natural evolutionary processes— randomly shuffling genes together and selecting for the hybrid combinations that work, rather than making just a few carefully designed constructs like previous studies—and showed we could indeed make new molecules without having to alter the region of the assembly line that houses the hypothetical proofreading machinery."
To make sure the researchers were not just working with unusually tolerant enzymes, they looked at the original model NRPS system from which the "proofreading" hypothesis was generated, he says.
"By targeting our preferred points in the assembly line, we showed we could make new molecules that should not be possible if there actually was a proofreading mechanism."
Their findings came as a "massive surprise," Professor Ackerley says.
"We had embarked on this study as believers that there was a proofreading mechanism, to try and hunt down exactly which parts of the assembly line had that role. The beauty of mimicking evolutionary approaches is that it doesn't matter how wrong your original ideas are–as long as the experiment is designed correctly, then you will arrive at the 'fittest' solution, even if it was totally unexpected."
More information: Mark J. Calcott et al. Efficient rational modification of non-ribosomal peptides by adenylation domain substitution, Nature Communications (2020). DOI: 10.1038/s41467-020-18365-0
Journal information: Nature Communications
Provided by Victoria University of Wellington