Mitochondrial metabolite mediates longevity through epigenomes
In a study published in Science Advances, researchers from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences revealed that mitochondrial metabolite acetyl-CoA links mitochondrial stress to the nuclear epigenome via NuRD complex for life-span regulation in C. elegans.
Metabolic homeostasis and aging are intimately linked. A regulatory center for cellular metabolism lies in the mitochondria. In addition to generating the bulk of adenosine 5′-triphosphate (ATP), mitochondria also generate many molecules such as lipids, heme, and intermediate metabolites that continuous communicate with the rest of the cell, allowing cells to integrate nutrient availability and energy demand to ensure cellular metabolic homeostasis.
Work in C. elegans has shown that mitochondrial stress during early life induces extensive chromatin restructuring that is essential for activation of the mitochondrial unfolded protein response (UPRmt), a process that promotes the recovery of mitochondrial protein homeostasis and stress-induced longevity.
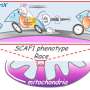
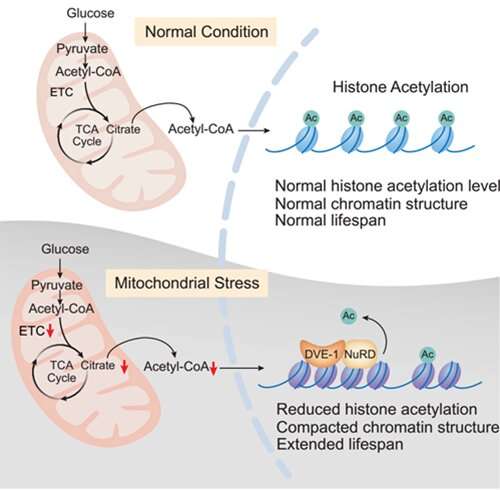
The researchers identified the NuRD complex that mediate the nuclear accumulation of the UPRmt transcription factor DVE-1 in response to mitochondrial stress. They further found that the impaired tricarboxylic acid (TCA) cycle upon mitochondrial stress results in a decreased level of citrate, which accounts for reduced production of acetyl-CoA and consequently induces nuclear accumulation of the NuRD and DVE-1, thereby enabling decreased histone acetylation and chromatin reorganization.
Restoration of the acetyl-CoA level by providing substrates and nutrients required for acetyl-CoA production is sufficient to counteract the chromatin changes and diminish the longevity upon mitochondrial stress.
Their findings uncover a novel molecular mechanism of the metabolite-mediated epigenome for the regulation of organismal aging.
More information: Di Zhu et al. NuRD mediates mitochondrial stress–induced longevity via chromatin remodeling in response to acetyl-CoA level, Science Advances (2020). DOI: 10.1126/sciadv.abb2529
Journal information: Science Advances
Provided by Chinese Academy of Sciences