Researchers reveal molecular basis for assembly of RuBisCO assisted by chaperone Raf1
A team led by Prof. Zhou Congzhao and Prof. Chen Yuxing from University of Science and Technology of China (USTC) of the Chinese Academy of Sciences (CAS) reported the crystal structures of Raf1 from cyanobacteria Anabaena sp. PCC 7120 and its complex with RuBisCO large subunit RbcL, and proposed a putative model for the assembly of cyanobacterial RuBisCO coordinated by the chaperone Raf1. The results were published in Nature Plants.
Photosynthesis is a fundamental biological process on Earth that provides the energy and sugars for most living organisms. In this process, ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO), which is the most abundant enzyme in nature, is responsible for converting the atmospheric CO2 into organic carbon. The folding and assembly of RuBisCO needs a series of chaperones, including the RuBisCO accumulation factor Raf1, which is highly conserved in cyanobacteria and plants.
The RuBisCO enzymes are generally classified into three forms. Form I, which is the most common form widespread in plants, algae, cyanobacteria and proteobacteria, is composed of eight large (RbcL, ~53 kDa) and eight small (RbcS, ~15 kDa) subunits, whereas form II and form III RuBisCO consist of only one or more RbcL dimers. Notably, RuBisCO is considered to be inefficient owing to its slow catalytic rate and unavoidable inhibition by O2, which impairs its CO2/O2 specificity. Thus, engineering RuBisCO to improve the carboxylation efficiency and CO2/O2 specificity is viewed as a promising strategy to increase crop growth and yield.
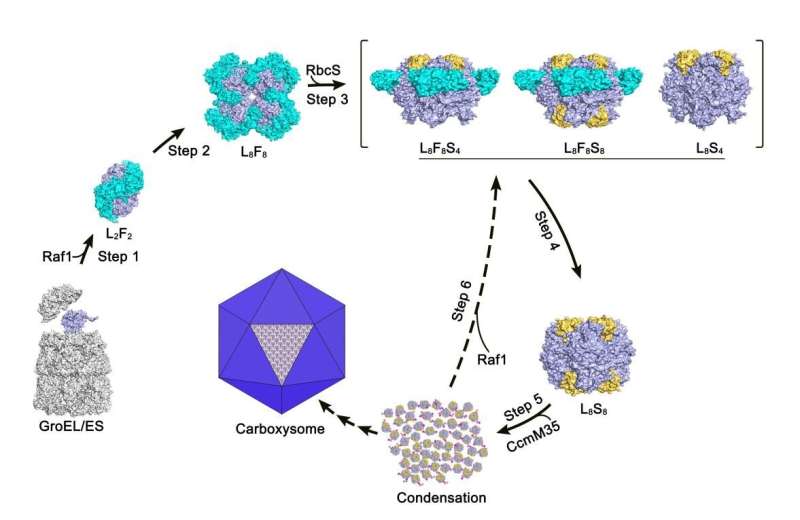
In the study, through structural analysis and biochemical assays, the researchers revealed that each Raf1 dimer captures an RbcL dimer, with the C-terminal tail inserting into the catalytic pocket, and further mediates the assembly of RbcL dimers to form the octameric core of RuBisCO.
Furthermore, they found that the cryo-electron microscopy structures of the RbcL–Raf1–RbcS assembly intermediates enabled them to see a dynamic assembly process from RbcL8Raf18 to the holoenzyme RbcL8RbcS8. In vitro assays also indicated that Raf1 can attenuate and reverse CcmM-mediated cyanobacterial RuBisCO condensation.
In all, the researchers presented the crystal structures of the full-length Raf1 and its complex with RbcL, in addition to the cryo-electron microscopy (cryo-EM) structures of a series of assembly intermediates that eventually form the RuBisCO holoenzyme.
Structural analysis combined with biochemical assays enabled them to figure out the fine molecular mechanism of Raf1-assisted cyanobacterial RuBisCO assembly.
More information: Ling-Yun Xia et al. Molecular basis for the assembly of RuBisCO assisted by the chaperone Raf1, Nature Plants (2020). DOI: 10.1038/s41477-020-0665-8
Journal information: Nature Plants
Provided by Chinese Academy of Sciences