Origins of genetic variability in seals
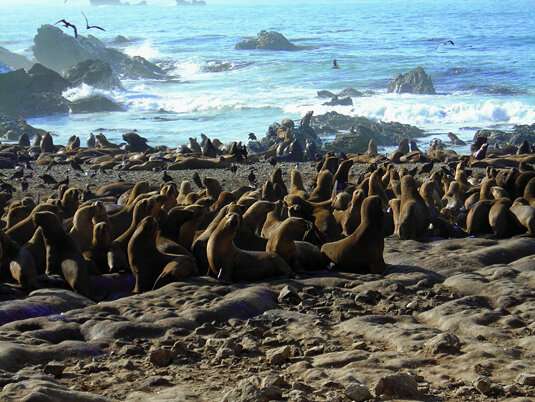

A new study led by Ludwig-Maximilians-Universitaet (LMU) in Munich researchers shows that fluctuations in population sizes in the past have had a significant effect on contemporary seal populations, and estimates the risk of genetic impoverishment in the species investigated.
In the course of Earth's history, evolution has given rise to an enormous range of biological diversity, which in turn enabled the emergence of complex, species-rich ecosystems. The availability of adequate levels of genetic variation is a basic prerequisite for evolution. Higher levels of genetic variability therefore increase the probability that any given population will be able to adapt to new environmental conditions and remain evolutionarily flexible.
Scientists led by LMU evolutionary biologist Jochen Wolf have examined the genetic variability of multiple seal species and show that a large part of today's variation is due to historical fluctuations in population sizes. In addition, the authors use the results of their genomic analyses to derive a parameter that allows them to assess the risk that genetic impoverishment and inbreeding pose to seal populations today. The new study appears in the journal Nature Ecology & Evolution.
Genetic variation is the product of random mutations, which are passed down from generation to generation. However, mutations can also be lost, owing to the effects of 'genetic bottlenecks', for instance. Such bottlenecks can occur when a large fraction of the population is lost. "It is generally assumed that populations that are made up of many individuals are likely to exhibit high levels of genetic variability," says Wolf. "We have now tested this assumption for 17 species of seals, by analyzing the genetic differences between 458 animals from 36 populations."
Since the genetic variation found in present-day populations can tell us a great deal about the genetic make-up of their ancestors, the authors of the study were able to deduce from their data how different populations have changed with time. "Genetic data are like a microscope that allows us to peer into the past," says Wolf. "The greater the differences between the genomic sequences, the farther back in time their last common ancestor lived. Our analyses enable us to look back thousands and even millions of years, and we can see that many populations must have gone through very narrow genetic bottlenecks—in other words, were drastically reduced in size—while others experienced significant expansions."
The researchers use the 'effective population size' as a measure of the extent of genetic variation within a population. This parameter is defined as the number of individuals that, under theoretically ideal conditions, would be expected to exhibit the same level of genetic variance as the real population of interest. The effective population size is related to, but much smaller than, the actual size of the real population, because the parameter includes the effects of factors such as reproductive behavior. Male seals in some species compete aggressively for females. That implies that the less dominant males may have no chance to reproduce, which in turn reduces the range of genetic variation in the following generation. "We assessed the impact of such effects, but our analyses indicate that the amounts of genetic variation in modern seals have been influenced mainly by historical fluctuations in population sizes, which are probably related to changes in the climate," says Wolf.
The ratio of the effective to the actual population size is often used to infer whether or not a given population possesses enough genetic variability to survive in the longer term. A very low quotient serves as a warning signal, since populations with low levels of variation are especially susceptible to inbreeding effects which, among other things, increase the risk of disease. "Most genetic studies undertaken in the context of conservation assess the level of genetic variability only across a few generations," says Wolf. "Our investigation, on the other hand, extends much further back in time. So we were able to take fluctuations in population sizes into account, and could calculate the population sizes we would expect to find today due to the genetic variability."
The expected population sizes were then compared with their actual sizes by means of a complex statistical procedure, which reveals whether the extant population is larger or smaller than the expected value. "This then tells us if a population is at risk because its current size is much too small to sustain that particular species in the longer term," says Wolf. In this context, the absolute number of individuals can be misleading. For instance, only 400 Saimaa ringed seals survive in the wild, and the species is regarded as endangered."
From a genetic point of view, however, despite their small number, we do not expect them to run into problems in the near future, as the animals are highly variable," says Wolf. The indications are that they settled in their present habitat only a short time ago—in evolutionary terms—and they retain the full range of variation that characterized their ancestors.
The situation in the Galapagos is quite different. There too, seal and sea lion populations are small, but their levels of genetic variability are also low—a factor which is not reflected in the value of the conventional ratio of effective to actual population size. The study shows that comparative genomic analyses of animal populations constitute an important tool for the identification of vulnerable populations in order to take protective measures.
More information: Claire R. Peart et al, Determinants of genetic variation across eco-evolutionary scales in pinnipeds, Nature Ecology & Evolution (2020). DOI: 10.1038/s41559-020-1215-5
Journal information: Nature Ecology & Evolution
Provided by Ludwig Maximilian University of Munich