How our single-celled relatives package their DNA
A group of single-celled organisms organises its DNA in a similar way to higher organisms such as plants, animals, and fungi. However, the way packaged DNA is read out differs between the two related groups, Bram Henneman discovered. Ph.D. defence on 5 December.
The most basic organisms on Earth are single-celled and have no cell nucleus. It was discovered around 1970 that these organisms are actually two different groups: bacteria and archaea. On the surface, they resemble each other. Their biochemistry, however, is fundamentally different, and therefore they are now considered to be two domains of life. The third domain includes organisms with cells that have a nucleus, the eukaryotes, which include humans.
Keep DNA safe and tidy
Archaea are closer to eukaryotes than to bacteria, their biochemistry shows. On this, the idea is based that eukaryotes are descended from an archaea ancestor. Recently, that theory is getting more and more popular. Now also the organisation of the DNA appears to be in line with it.
DNA molecules are extremely thin and long. To keep them safe and tidy, cells twist DNA strands tightly together, making them compact. At the same time, genes that are on DNA molecules must be accessible to the cell parts that read them and copy them into messenger RNA, when they need to be expressed. The so-called DNA-binding proteins are responsible for the organisation of the DNA (so making it compact, regulating expression).
"There are different types of these proteins," says Henneman. "I have dedicated myself to one type, the histones. They occur in archaea and eukaryotes, but not in bacteria. They can shed light on the evolution of eukaryotes."
Beads and rods
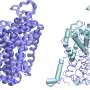
Histones of archaea appear to function slightly differently than those of eukaryotes. With eukaryotes, eight histones form a bead around which a piece of DNA is wrapped twice. These beads are called nucleosomes; wrapped beads lie side by side like a chain.
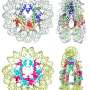
In archaea, such beads are not found, but the histones organise the DNA in a different way. Some archaea histones form units of two pieces, around which a piece of DNA bends. Henneman: "An indefinite number of those units is piled up into a hypernucleosome, as we have called it. That is a rod-shaped structure around which a long piece of DNA is wrapped."
With the nucleosomes of eukaryotes, chemical groups on the histones control whether or not a piece of DNA is read. The hypernucleosomes of archaea are probably not accessible for transcription into RNA, Henneman thinks. Yet the DNA is not permanently stored in hypernucleosomes, he explains. "Because units can be attached to and removed from such a hypernucleosome, it is a dynamic whole of variable length. Depending on the circumstances, more or less DNA is protected against reading."
Salt crust
"The thing I liked most about my research was that during it, a new archaea species was found that lives on salt crusts in the desert, a Nanosalina species. We were the first to study the histones of this new species. It is not cultivable, but we were able to sequence the DNA encoding three of its histone sequences, replicate it and express it in a bacterium to see what they did with DNA. Based on the DNA sequence and our previous research, we expected that one of the three histones we examined could form a hypernucleosome. That seems to be correct."
Bram Henneman defended his thesis Histone-DNA assemblies in archaea—shaping the genome on the edge of life on 5 December.
Provided by Leiden University