Scientists identify harmful bacteria based on its DNA at a very low cost

A new bacterial identification method, called ON-rep-seq, examines selective, strain-specific fragments of the bacterial genome, allowing the generation of results that earlier required DNA sequencing of the entire bacterial genome or tedious approaches like pulsed field gel electrophoresis, which previously has been the gold standard for strain-level typing of microorganisms. Hence, the method has the potential to change the approach utilized for investigating food-based disease outbreaks by making analysis much less time- and cost consuming.
Today, bacterial detection and identification based on bacterial DNA requires expensive instrumentation and many hours of work by highly trained specialists. Let's imagine, for example, there is a suspected Salmonella outbreak. Usually in order to locate its origin, not only will investigators have to analyze many samples, but the analysis has to be precise in order to distinguish one bacterial strain from another.
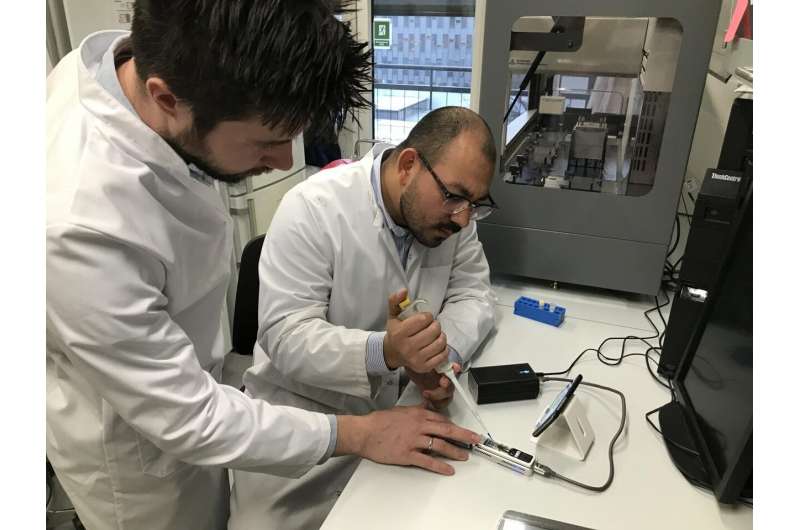
"Our new method allows identification and typing of hundreds of samples in less than two hours, and we expect that this will even be reduced to "real time" in a short period of time," says one of the researchers behind the study, Lukasz Krych, Associate Professor at the Department of Food Science at the University of Copenhagen, Denmark.
Method builds on a device that was used for DNA sequencing in space
The new method is based on nanopore sequencing, which is a new, real-time DNA sequencing approach "that will definitely revolutionize the future of DNA sequencing," according to Lukasz Krych.
The research project was carried out in collaboration with the polish company GenXone S.A., which helped to set up a bioinformatics pipeline that is needed to perform fast and efficient analysis of the sequencing data.

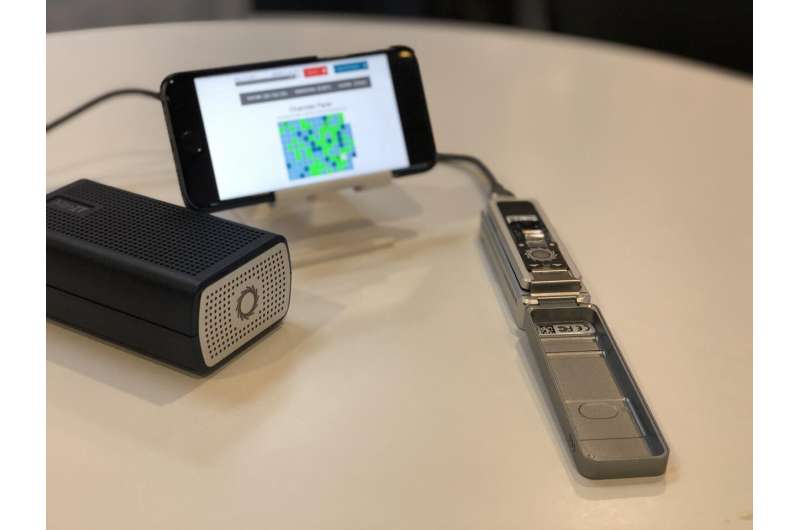
The smallest ever sequencer offered by Oxford Nanopore Technologies, called MinION, is a $999 hand-held, USB-powered device that became commercially available in 2015. A year later it was taken to the International Space Station, where it achieved the first DNA sequencing in history performed in zero-gravity conditions. Despite the indisputable revolution in DNA sequencing offered by MinION, it quickly became clear that the data generated with the device are still not perfect due to sequencing errors, for example, while the analysis remained relatively expensive to perform (approximately $150 per bacterium).
Small device with fast and cheap analysis offers huge opportunities within food safety
The scientists from the Department of Food Science at the University of Copenhagen have found a way to utilize this technology to analyze hundreds of bacteria at a time, cutting costs to less than $2 per bacterium, while at the same time increasing the accuracy to more than 99%.
"Our method can be used both within food safety organizations, where you can quickly find disease-causing or health-promoting bacteria, and also in the health sector, where you will be able to perform certain analyses that are not even considered today because of the price and time-consuming nature of traditional analysis," says another of the researchers behind the study, Postdoc Josue Leonardo Castro Mejia.
At the moment, there are several companies testing the method to implement in their systems for establishing rapid screening programmes for thousands of strains.
More information: Łukasz Krych et al, DNA enrichment and tagmentation method for species-level identification and strain-level differentiation using ON-rep-seq, Communications Biology (2019). DOI: 10.1038/s42003-019-0617-x
Journal information: Communications Biology
Provided by University of Copenhagen