Solving a combinatorial quandary

Why do certain proteins in the body bind with some substances, but not with others?
The answer could be the difference between a drug working or not. The answer, however, is elusive by virtue of the sheer scope of mutations that make proteins vary between each other, and between individuals.
"Amino acids are the building blocks of proteins, and my copy of a protein might have just one that is different than yours," says Brian Chen, an associate professor of computer science and engineering at Lehigh University's P.C. Rossin College of Engineering and Applied Science. "But there are many different ways that one amino acid may change, and there are many different amino acids that may potentially change. So the natural question is, why don't you test all the possibilities? Well, that would be a combinatorial nightmare. There are just too many possibilities to test in a wet lab, too many possibilities for a human to simulate on a computer, and too many possibilities for a person to keep straight in their head and consider in a systematic way."
Currently, researchers must review the data and do their best to interpret whether or not proteins are interacting, and how.
Supported by a four-year nearly $1 million grant recently awarded by the National Institutes of Health, Chen is developing software that can both handle the scale of possibilities and replace human interpretation of that data. But in a novel twist, the software will also generate an English language translation that will explain the mechanism behind protein interaction or noninteraction. In essence, it will tell the researcher what it thinks is happening.
"The software provides the mutations to investigate and the reasons why it thinks those mutations are significant," says Chen. "And that tells you something about the experiment you might do. You could take a carpet-bombing approach where you spend a lot of money and test all these different combinations of mutations, and to some extent, a lot of science involves something like that. But this software allows us to reduce the scope of testing because it gives a rationale for that testing."
The software will eventually utilize four of the biophysical mechanisms that control protein interactions—shape complementarity, electrostatic complementarity, hydrogen bonding, and hydrophobicity. It currently uses the first two, and—thanks to a collaboration with Rutgers University—has already successfully demonstrated the ability to predict something completely unknown.
In that partnership, supported by the National Science Foundation and the NIH, Chen worked with Nilgun Tumer. Tumer is a plant pathologist at Rutgers, whose lab is studying ricin, a poison found naturally in castor beans.
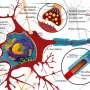
"Ricin is a toxic agent that has unfortunately been weaponized," says Chen. "It binds to the ribosome, which is this big machine in your cells that makes proteins, and breaks it. Your body constantly requires proteins to do essential tasks, like clot blood, and that's why ricin is so deadly. Right now, there's nothing that prevents the toxin from getting to the ribosome. So there's no treatment for ricin exposure, and if you're exposed to enough of it, you'll die."
Tumer's group is studying the interaction between the toxin and the ribosome. Specifically, they're looking at how the ricin recognizes and binds to a specific protein called the P-protein, which is located on a stalk of proteins within the ribosome.
"So, again, that's an interaction where specific amino acids are important for selecting where the ricin will land, and how it will land," Chen explains. "Tumer and her group had a theory and a model we could build, and we made some predictions. It turned out that some of the amino acids predicted by our software hadn't been considered before in the interaction. They tested these amino acids, and found that when they were substituted, it was harder for the ricin to bind, causing it to be less toxic. In the past, the software had predicted things that were known, but that was the first time it predicted something completely unknown. We were able to add something new to that study."
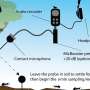
Chen's NIH grant, "Algorithmic Identification of Binding Specificity Mechanisms in Proteins," will support the continued development and testing of the software, the latter of which will be done in collaboration with Julie Miwa, an associate professor of neuroscience in the Department of Biological Sciences in Lehigh's College of Arts and Sciences. Miwa's lab studies the interaction between lynx proteins and nicotinic acetylcholine receptors in the brain.
"One of the questions that comes up in her research is, do certain variations of these lynx proteins turn the receptors on or off? And that's really important because it's believed that lynx mutations affect neuroplasticity. Discovering new mutations that affect this interaction could shed light on the mechanisms that affect learning and anxiety. But there are hundreds of possibilities. So I'll be using my software to suggest mutations that could interrupt binding between the lynx and acetylcholine receptors. It's a great opportunity to validate the software from our lab, because it's a completely blind scenario, and a great opportunity to assist Dr. Miwa's study of the lynx family of proteins. We don't know which amino acids are going to be important, but her team will reveal if we were right or wrong when they find the biological truth."
The ultimate goal of this research, he says, is for the software to utilize more than four biophysical mechanisms in its predictions and to provide even more robust explanations for why mutations might affect binding. He also looks forward to more opportunities for collaboration, especially with students, in this work, which he calls "super exciting" and "deeply fascinating."
"The interest in research among undergraduates here at Lehigh is really high," says Chen, "and it's allowed me to get a lot of research done on this specific project that I otherwise would not have been able to do, specifically in terms of evidence and validation. I think undergraduate research is one of the great strengths of this university."
Provided by Lehigh University