Understanding transporter proteins at a single-molecule level

Like a boat helping passengers cross a river, transporters move substances across cell membranes. This process is fundamental to the healthy functioning of cells in life forms from bacteria to humans. The function of these transporters previously had to be inferred from the behavior of hundreds or thousands of them working together. Published today in Nature, new techniques enable the study of one transporter at a time.
This powerful advance comes into focus if you imagine trying to understand the location, speed and passengers of a particular boat given only the combined statistics for the entire fleet.
"By looking at the activity of single molecules, we have clarified part of the mechanism underlying transporter activity that may prove critical for future research on the many clinically relevant proteins in this family," said co-corresponding author Scott Blanchard, Ph.D., a member of the St. Jude Department of Structural Biology.
Technique opens new possibilities
This research relied on a technique called single-molecule fluorescence resonance energy transfer or smFRET. The method enables investigators to gather precise measurements of the activity of individual transporters. This leading-edge technique is a powerful tool for studying mechanisms of action as well as mutations implicated in disease. It is available in a handful of laboratories worldwide.
Previous single-molecule techniques had the capacity to measure the the activities of so-called "ion channels" that allow charged particles to rapidly cross cell membranes.
These single-molecule methods revolutionized understanding of channels but were not amenable to transporters due both to the wide variety of transported substances and their relatively slow rates of transport. The new smFRET methods are more useful for a wide variety of transporters.
Transporters, step-by-step
Neurotransmitter: sodium symporters (NSS) are a family of transporter proteins, found most notably in the brain, that move molecules into and out of cells. In humans, NSS for the neurotransmitters serotonin and norepinephrine are the targets for nearly all antidepressants. NSS for dopamine are the key targets for amphetamine and cocaine. Understanding this class of proteins and how they function will give novel insights into the mechanisms of these therapeutic and abused drugs and how therapies targeting these transporters can be improved.
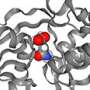
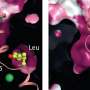
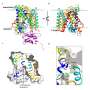
The researchers applied smFRET to study a bacterial relative of NSS proteins: the MhsT transporter, which transports amino acids across the cell membrane. The team wanted to understand the slowest part of the transport process, called the rate-limiting step. The researchers were surprised to learn that the rate-limiting step for the MhsT transporter is different for different cargo.
To move molecules through a membrane, transporters change their shape, both to pick up substances on the outside of the cell and to release the substances on the inside. The researchers revealed that the slowest part of the cycle is the process of changing shape back to the outside, when the transporter was thought to be emptiy of some cargoes but not others.
"Since different cargoes have different slow steps, the findings suggest that the 'return' step is not empty, but rather fits with other evidence of a secondary binding site on the transporter protein that is key to regulating its activity," said co-corresponding author Jonathan Javitch, M.D., Ph.D., of Columbia University and the New York State Psychiatric Institute.
A better understanding of the functional relevance of this second binding site within the NSS family of proteins will be critical to studying the pharmacology and efficacy of drugs targeting these transporters.
More information: Quantifying secondary transport at singlemolecule resolution, Nature (2019). DOI: 10.1038/s41586-019-1747-5 , nature.com/articles/s41586-019-1747-5
Journal information: Nature
Provided by St. Jude Children's Research Hospital