A catalog of DNA replication proteins

Maintenance of genome integrity—and prevention of diseases such as cancer—requires complete and faithful replication of the genome every cell division cycle.
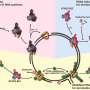
To fully understand how genome integrity is maintained, David Cortez, Ph.D., and colleagues have generated a "catalog" of the proteins present at sites of DNA duplication (replication forks) and chromatin packaging of newly synthesized DNA.
The investigators used a method they developed called iPOND (isolation of proteins on nascent DNA) combined with quantitative mass spectrometry in multiple cell types to identify 593 proteins that are enriched at replication forks and nascent chromatin. Using loss-of-function genetic analyses and a review of existing studies, they found that 85% of the proteins had activities consistent with a function in DNA and chromatin replication or replication stress responses.
They demonstrated the value of their catalog by identifying a role for BET family proteins in controlling DNA replication. The resource was published in the Sept. 24 issue of Cell Reports.
More information: Sarah R. Wessel et al. Functional Analysis of the Replication Fork Proteome Identifies BET Proteins as PCNA Regulators, Cell Reports (2019). DOI: 10.1016/j.celrep.2019.08.051
Journal information: Cell Reports
Provided by Vanderbilt University