Unraveling the three-dimensional genomic structure of male germ cells
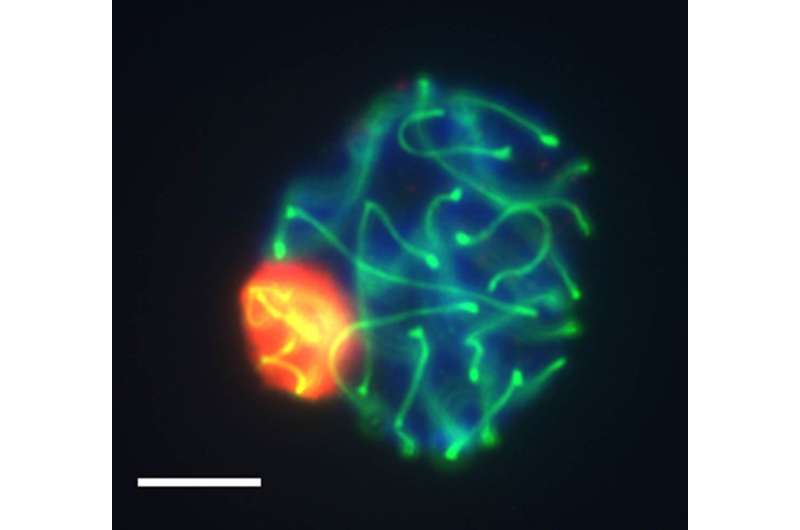
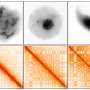
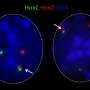
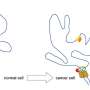
The genome is not just a linear sequence of letters, but is also compartmentalized into a specifically tailored chromatin structure within cell nuclei. This three-dimensional genomic structure is fundamental, given that it determines which genes are turned on and off in each cell type.
A new study led by scientists from the UAB and the CNAG-CRG and published in Cell Reports reveals the three-dimensional genomic structure of male germ cells. The study, carried out on mice, shows that this structure is extremely dynamic during the formation of germ cells (gamete precursor cells). Moreover, the study revealed a fine-tuned balance between chromatin remodeling, architectural proteins such as cohesins and gene expression during this process.
All sexually reproducing organisms form haploid gametes (oocytes and sperm), each cell type carrying only one copy of each chromosome, through two consecutive cell divisions preceded by one round of genome replication. This process is known as meiosis, and implies that the genome must be packaged and unpackaged in a precise and highly regulated manner.
"Our work shows the dynamics of chromatin remodeling during the formation of male gametes by comparing changes in chromatin folding and gene transcription at different moments throughout male meiosis," says coordinator of the study Aurora Ruiz-Herrera, researcher at the Department of Cell Biology, Physiology and Immunology of the Institute of Biotechnology and Biomedicine (IBB) at the UAB, where she leads the research group in Animal Genomics.
"We have thus demonstrated the existence of different degrees of genome folding and how these different levels of genome organization are related to structural proteins such as cohesins and gene expression. The results will pave the way for new investigations into the molecular mechanisms regulating these changes."
"This study has been possible only thanks to a combination of complementary techniques in biology such as molecular genetics, microscope imaging and computer simulations. It truly is a multidisciplinary project," explains Marc A. Marti-Renom, ICREA researcher and head of the Structural Genomics Group at the CNAG-CRG and co-leader of the study.
The project represents a significant advance in the study of the mechanisms generating and regulating the 3-D structure and function of the genome during the formation of gametes. Determining these mechanisms is fundamental, given that the deregulation of this process can lead to diseases such as infertility and chromosome alterations like trisomy 21.
According to scientists, the research also represents an example of the importance of synergy among specialists from different fields such as molecular and cell biology, genomics and bioinformatics in advancing in our knowledge of the regulation and structure of the genome.
More information: Covadonga Vara et al. Three-Dimensional Genomic Structure and Cohesin Occupancy Correlate with Transcriptional Activity during Spermatogenesis, Cell Reports (2019). DOI: 10.1016/j.celrep.2019.06.037
Journal information: Cell Reports
Provided by Autonomous University of Barcelona