Methods in belowground botany
Plant root systems play a crucial role in ecosystems, radically impacting everything from nutrient cycling to species composition. Despite their importance, scientists are just beginning to develop the tools to understand how these complex systems are structured, how they function, and how structure and function are related. Much of the research into root systems today uses sophisticated new technologies to address basic questions and descriptions. A recent special issue of Applications in Plant Sciences contains six papers from the cutting edge of root science, exploring questions from the subcellular to the ecosystem level.
Because research into root systems is relatively new, much research is still in the realm of description. For example, Kengkanna et al. (2019) evaluate root structure in cassava, refining which variables might be agronomically important and could thus serve as a target for breeding programs. They do this using a tool called digital imaging of root traits (DIRT), a high-throughput method for scoring root phenotypes that is itself only five years old. "In areas where drought is or already has become more prominent, [DIRT] makes it feasible for breeding programs to target root traits that may be critical to achieving higher yield with limited water supply," said Dr. Gregory Pec, a postdoctoral research associate at the University of New Hampshire, and co-editor of this special issue.
Other current root system research takes aim at basic, fundamental questions, such as: are the microbes growing under a particular tree species better adapted to decompose that species' leaf litter? This idea, known as the home-field advantage hypothesis, was tested by Martini et al. (2019); the authors found that the answer might be a little more complicated, depending on fine-scale species composition, and which nutrient is under investigation.
Metzler et al. (2019) address the question: given a mixed sample of soil containing many roots, how can one figure out which roots belong to which species efficiently? The authors use fluorescent amplified fragment length polymorphisms—-fluorescently marked small segments of DNA—-to determine species identities of multiple roots in mixed samples simultaneously, dramatically cutting cost and time over previous Sanger sequencing methods. In the process, they provide root size profiles for 193 species, effectively doubling the existing database. Genetic tools have moved root system research forward on many fronts, aside from identifying which roots belongs to which species.
One area of particular promise for genetic research is in understanding and mapping soil microbial diversity. "The use of traditional, cultivation-based approaches to examine soil-microbial-root dynamics has proven challenging as these methods retrieve a small fraction of the total diversity. However, technological advances in the last decade have allowed for these complex and often context dependent interactions to be investigated," said Dr. Pec.
In this issue, a historical dimension is added to this genetic research into root-microbe associations, as Heberling and Burke (2019) present research to investigate and quantify arbuscular mycorrhizal fungal communities on historical herbarium specimens. This reconstruction of historical plant-microbe interactions can help us understand how climate change has impacted forest ecosystems.
Research into root systems was neglected for decades due to the difficulty of studying these delicate, complex, underground systems. Today, as improvements in technology have made this research much more feasible, interest in root systems is growing, and databases are starting to fill with important baseline data like root structure and size profiles, and microbial associations. "Our understanding of root structure-function relationships is often limited due to methodological challenges associated with roots' hidden nature in soils," said Dr. Pec. "In the 1990s, plant ecologists started looking at belowground competition and root biomass distributions, but this did not require new methodology. In the 2000s, mycorrhizae, plant-soil feedback and microbial interactions of root ecology really took off. However, prior to that, most research by mainstream plant scientists was just not focused on roots."
That is beginning to change.
More information: Gregory J. Pec et al, Methods in belowground botany, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1239
Susan Fawcett et al. Tracking microhabitat temperature variation with iB utton data loggers, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1237
J. Mason Heberling et al. Utilizing herbarium specimens to quantify historical mycorrhizal communities, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1223
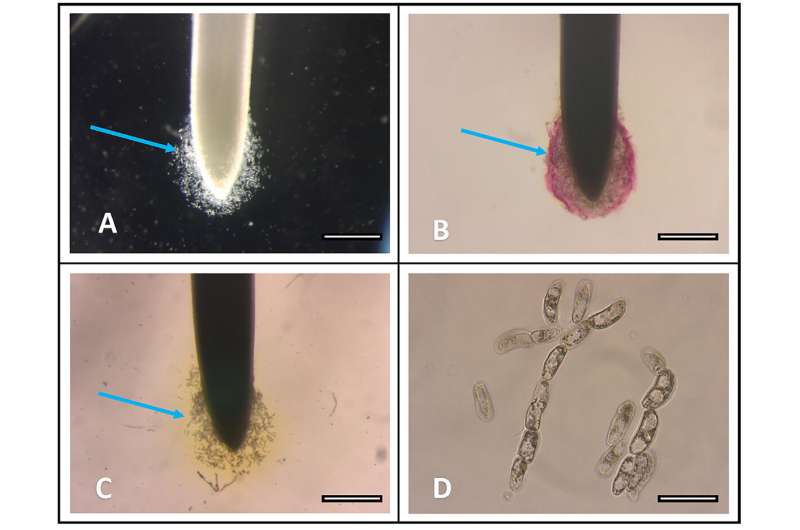
David A. Huskey et al. Use of rhodizonic acid for rapid detection of root border cell trapping of lead and reversal of trapping with DN ase, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1240
Jitrana Kengkanna et al. Phenotypic variation of cassava root traits and their responses to drought, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1238
Francesco Martini et al. Small‐scale and multi‐species approaches for assessing litter decomposition and soil dynamics in high‐diversity forests, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1241
Paul Metzler et al. Expanding and testing fluorescent amplified fragment length polymorphisms for identifying roots of boreal forest plant species, Applications in Plant Sciences (2019). DOI: 10.1002/aps3.1236
Journal information: Applications in Plant Sciences
Provided by Botanical Society of America