Meet B. fragilis, a bacterium that moves into your gut and evolves to make itself at home
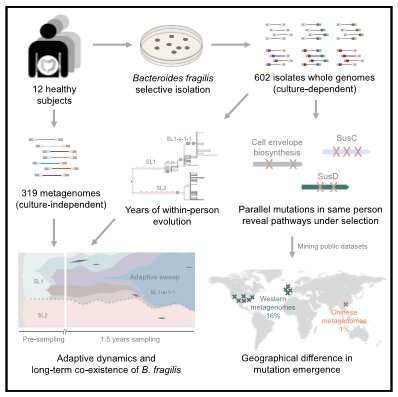
MIT researchers have analyzed population genomics and metagenomics to investigate the microbiome evolution of Bacteroides fragilis, one of the most prevalent bacteria found in humans' large intestines. In a paper published April 23 in the journal Cell Host & Microbe, the authors describe how the common gut microbe adapts and evolves within individuals as well as across Western versus Eastern cultures.
"The strains of B. fragilis that are growing in humans have been in that gut-like environment for millions of years, so the idea that encountering a new host's gut would induce a bunch of new adaptive mutations, and that these commensals would still be rapidly evolving, was surprising to us," says Eric Alm, a professor of biological engineering and co-director of the Center for Microbiome Informatics and Therapeutics at MIT.
Alm and his colleagues' analysis revealed at least sixteen genes undergo within-person evolution, and the majority of mutations targeted pathways involved with fiber uptake in the cell and biosynthesis of the cell envelope. While the mutations they observed occurred in the same types of genes over and over, and within similar locations in those genes, the particular gene that acquired mutations in each person was often different, suggesting ongoing personalization of the bacteria. "Bacteroides species are helping digest complex fibers in the large intestine, which are coming from the food you eat, so adaptations in them may be related to the personalized diet," Shijie Zhao, co-first author and member of Alm's lab, says.
The explanation as to why those genes are being targeted may not be limited to dietary choices alone, however. For example, it is possible that rapid evolution is a tactic needed to avoid phages and immune cells. "These mutations might be responsible for changing the way B. fragilis interacts with the immune system," says Tami Lieberman, co-first author and now an assistant professor in MIT's Department of Civil and Environmental Engineering. "Alternately, it could be a technique to enhance their ability to fend off attacks from other members of the microbiome."
Additionally, the authors found differences in mutations between Western and Eastern metagenomes, with one allele highly selected for within Western participants—with independent mutation in each person—but rare in Chinese subjects. This striking difference is in a gene with an unknown biological role, but the authors believe that figuring out the source of different pressures across continents may yield new discoveries.
Typically, when researchers consider changes in the microbiome, they are looking to see if the species composition of the microbiome is remaining constant, not what might be changing within one particular species. But in light of these findings, keeping an eye on the adaptive abilities of individual commensals, particularly as it concerns probiotic choices and microbial therapies, could become more of a focus.
"It's not a scary thing—I think we just need to be aware of and thoughtful about the fact that you can do extensive testing in a lab, but once a commensal gets out there and into a real person in the real world, it can evolve—its phenotype can change—and that's something that might be hard to anticipate," Alm says.
Moving forward, the authors plan to undertake more mechanistic studies and conduct metagenomic and genomic analyses in different species of commensals, under both disease and healthy conditions, to better understand the extent of evolution within the microbiome.
"If this pattern of gut bacteria evolving really quickly after introduction into a new host holds up, then we'll probably see entirely different pathways in different types of bacteria, and that's going to be pretty exciting," says Alm.
More information: Shijie Zhao et al, Adaptive Evolution within Gut Microbiomes of Healthy People, Cell Host & Microbe (2019). DOI: 10.1016/j.chom.2019.03.007
Journal information: Cell Host & Microbe
Provided by Cell Press