March 28, 2019 report
Researchers find associations between structural variation in gut microbiome and host health

A team of researchers from Israel, the U.S., the Netherlands and Norway has found what they describe as associations between structural variation in the human gut microbiome and host health. In their paper published in the journal Nature, the group describes their genetic study of microbes living in the guts of large groups of people from Israel and the Netherlands, and what they found.
In recent years, medical scientists have become increasingly focused on the microbes that live in and on our bodies with the goal of figuring out which are good or bad for us, and in what ways. In this new effort, the researchers sought to better understand gut bacteria that are genetically very similar. They wanted to know if such small differences are due to natural instances between species or if they simply represent the evolution of bacteria as they exist in the gut.
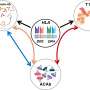
To learn more, the researchers obtained 887 microbiome samples from people living in Israel and studied them using iterative coverage-based read assignment and SGV-Finder algorithms. They report that they found 7,479 variants in 56 of the species they isolated in the samples. They further report that of those found, they were able to identify 5,056 deletions in genomes and 2,424 variable structure variants, which, they note, were prevalent in genes that were involved in CRISPR and antibiotic functions. Those involved in housekeeping, on the other hand, were noticeably less prevalent. They further report that the structural variants they looked at harbored genes with distinct functions that could be associated with bacterial growth—possibly suggesting utility in adaption. They also found 124 associations between variants and diseases in the host such as those connected to body mass index, blood pressure and cholesterol levels.
The researchers performed a similar analysis on microbiome samples obtained from 1000 people living in the Netherlands, and found many of the same structural variants with many of the same associations. They conclude by suggesting that their results support prior findings in studies of the mouse microbiome and that more research is needed to fully understand the role of genetically similar microbes living in our guts.
More information: David Zeevi et al. Structural variation in the gut microbiome associates with host health, Nature (2019). DOI: 10.1038/s41586-019-1065-y
Journal information: Nature
© 2019 Science X Network