July 14, 2016 report
Study reveals some secrets of how microbes and their genes traverse the globe

(Phys.org)—An international team of researchers led by MIT's Ilana Brito has discovered some of the ways that certain microbes are able to travel around the world. In their paper published in the journal Nature, the team describes how they collected samples of microbes from villagers living in Fiji to learn more about the nature of microbe transfer.
Human beings, as most are aware, are populated by microbes, both inside and out—but how these microbes are transferred between people is still something of a mystery. In this new effort, the team flew to Fiji to study the microbes of people living in remote villages—they were chosen because they are both remote and relatively stationary. This allowed the researchers to obtain samples of microbes from several parts of the body in people that are not likely to be getting or receiving new microbes as regularly as people living in more transient places, such as the United States.
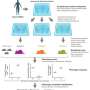
The team collected feces, saliva and swabs from the hands of 172 people living in five different villages (and from 81 people in North America). Their goal was to see if they could use an analysis of the microbes found to create a social map that showed the spread of microbes across a region. As part of their analysis, they sequenced individual microbe cells to see whole sets of genes, which was necessary for identification.
The team reports that they used some of their data to create a map and found that simple, everyday activities can have an impact on the transfer of microbe genes present in the human microbiome—even across species. As one example, they found that people living in Fiji tend to have more microbes in their guts than do people in North America—including one type in particular that helps digest starch. They were even able to see slight differences in microbe amounts between people in different villages, which offered a peek into the ways that such microbes can travel—such as through routine interactions like sharing a meal—an exchange that can alter a microbiome. Such changes, the team suggests, could explain why different groups of people can be more or less susceptible to different diseases, such as those that cause digestion problems, allergies or even asthma.
More information: I. L. Brito et al. Mobile genes in the human microbiome are structured from global to individual scales, Nature (2016). DOI: 10.1038/nature18927
Abstract
Recent work has underscored the importance of the microbiome in human health, and has largely attributed differences in phenotype to differences in the species present among individuals. However, mobile genes can confer profoundly different phenotypes on different strains of the same species. Little is known about the function and distribution of mobile genes in the human microbiome, and in particular whether the gene pool is globally homogenous or constrained by human population structure. Here, we investigate this question by comparing the mobile genes found in the microbiomes of 81 metropolitan North Americans with those of 172 agrarian Fiji islanders using a combination of single-cell genomics and metagenomics. We find large differences in mobile gene content between the Fijian and North American microbiomes, with functional variation that mirrors known dietary differences such as the excess of plant-based starch degradation genes found in Fijian individuals. Notably, we also observed differences between the mobile gene pools of neighbouring Fijian villages, even though microbiome composition across villages is similar. Finally, we observe high rates of recombination leading to individual-specific mobile elements, suggesting that the abundance of some genes may reflect environmental selection rather than dispersal limitation. Together, these data support the hypothesis that human activities and behaviours provide selective pressures that shape mobile gene pools, and that acquisition of mobile genes is important for colonizing specific human populations.
Journal information: Nature
© 2016 Phys.org