Researchers reveal active-state structure of popular drug target for blood pressure
Bringing a long quest to a satisfying conclusion, researchers have mapped the active-state structure of the angiotensin II type 1 receptor, the target of widely prescribed drugs to regulate blood pressure and kidney function.
The study, published online Jan. 10 in Cell, was conducted by researchers in the Blavatnik Institute at Harvard Medical School and colleagues at Duke University Medical Center.
When the hormone angiotensin attaches to this receptor, it constricts blood vessels to raise blood pressure and stimulates the adrenal glands to retain salt in the kidneys. An estimated 5 percent of adults in the United States take angiotensin II receptor blockers to lower blood pressure, treat heart failure or prevent kidney failure or stroke.
Despite the receptor's critical roles in the body and its popularity as a drug target, researchers have struggled to answer the question: How does it activate?
"Millions of people take drugs that turn this receptor off, and it has been studied very thoroughly, but because of technological limitations, people couldn't see what it looks like when it's turned on," said Andrew Kruse, associate professor of biological chemistry and molecular pharmacology at HMS and co-senior author of the paper with Robert Lefkowitz of Duke.
"Our study provides a framework for interpreting a lot of data that's been collected over the years about the receptor's function, helping to turn disparate pieces into a single coherent model," Kruse said.
The new findings also offer clues about how drugs might be developed that activate, rather than block, the angiotensin II receptor or other receptors in highly controlled ways.
Twist and shout
Like many proteins embedded in the cell membrane, the angiotensin II receptor twists into different shapes depending on whether it's inactive or bound to one protein versus another.
Researchers who want to study a receptor in a specific shape, or conformation, often try to generate an antibody with a complementary shape that will hold the receptor still. Then they can crystallize it, bombard it with X-rays and translate the resulting image into a 3-D atomic structure.
Lefkowitz's group tried the standard tactic—inoculating a llama to stimulate its immune system to produce the needed antibodies—but each attempt proved unsuccessful.
"The receptor would flip-flop between all these different conformations," said Lefkowitz. "We tried for at least six years to develop an antibody that would stabilize it, and we failed at every turn."
A tool developed by the Kruse lab in 2018 provided the key. Using yeast instead of llamas, the researchers had generated a library of 500 million artificial nanobodies (small antibodies) to help structural biologists.
One of the nanobodies did exactly what Lefkowitz needed.
"Andrew's new technique, combined with lots of hard, collaborative effort, finally made it possible," he said.
"This is an example of a good collaboration, where we each provided something unique," Kruse agreed. "We offered our nanobody library and expertise in crystallizing G protein-coupled receptors like the angiotensin II receptor, while the Lefkowitz lab brought their strengths in receptor biochemistry and pharmacology."
Conor McMahon, a postdoctoral researcher in the Kruse lab and co-first author of the paper, together with Laura Wingler and Dean Staus in the Lefkowitz lab, added, "This demonstrates the utility of our nanobody library, and it's particularly exciting to see it succeed in a case where llama immunization was unsuccessful."
Details revealed
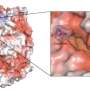
Capturing the receptor's intricately detailed crystal structure revealed how it binds to angiotensin, how changes on the portion of the receptor outside the cell triggers changes on the portion inside the cell, and more.
"Many of the conformational changes were unique compared to those we'd seen before," said Kruse.
Kruse and team also hope the study will help provide a path to answering questions about the mysterious phenomenon known as biased agonism, in which a protein that binds to a receptor activates one pathway preferentially, rather than activating two or more pathways equally.
Journal information: Cell
Provided by Harvard Medical School