Pushing microbes to deliver preferred products
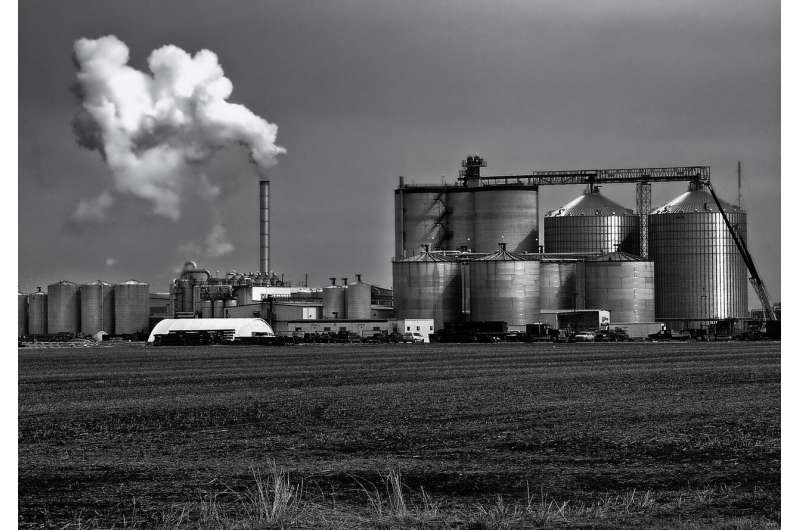

If environmental engineer Daniel Noguera had his way, he would orchestrate a microbiome to pump out higher-value chemical products.
In that vein, Noguera, the Wisconsin Distinguished Professor of civil and environmental engineering at the University of Wisconsin-Madison, along with graduate student Matt Scarborough and collaborators in the Great Lakes Bioenergy Research Center, recently published a paper in the journal mSystems in which they analyzed the makeup and metabolic activity of a mixed microbial community within a bioreactor. By doing so, they're edging closer toward engineering such a community to optimally generate particular products—an outcome that would boost the utility—and financial viability—of the residue of biofuel production.
In their work, the researchers studied the organic leftovers of lignocellulosic ethanol production with an eye toward optimizing the yield of medium-chain fatty acids, which are useful precursors for industrial chemicals and pharmaceuticals. The medium-chain fatty acids are an alternative product—with a higher potential financial value—than the methane that's typically generated from the residues of ethanol production.
"If we think about the residues from that process and generate more value from those residues, the hope is that we improve the economy of the overall system," says Noguera.
The researchers identified the members of the microbiome and their roles in the process, while using thermodynamic analysis to examine potential ways to drive production of the medium-chain fatty acids.
"We can establish communities that make these materials, but the ratio of medium-chain fatty acids to other microbial products is not optimized," says Noguera. "So how do you get the microbes to change? Or how do you engineer that community to make more of what you want? That's still an open-ended question."
More information: Matthew J. Scarborough et al. Metatranscriptomic and Thermodynamic Insights into Medium-Chain Fatty Acid Production Using an Anaerobic Microbiome, mSystems (2018). DOI: 10.1128/mSystems.00221-18
Provided by University of Wisconsin-Madison