Himalayan marmot genome offers clues to life at extremely high altitudes
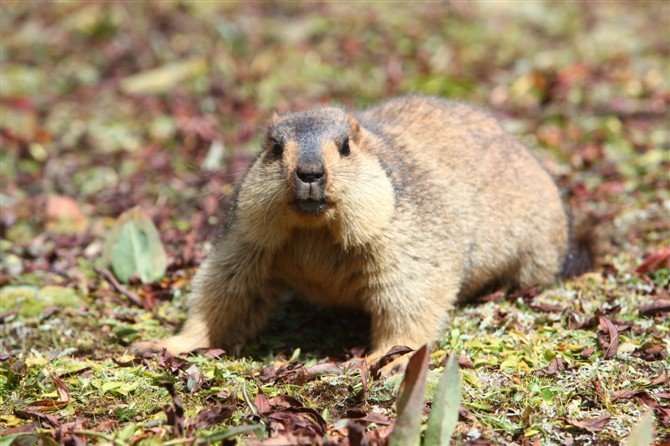

Himalayan marmots can survive at altitudes up to 5,000 meters in the Himalayan regions of India, Nepal, and Pakistan and on the Qinghai-Tibetan Plateau of China, where many of them face extreme cold, little oxygen, and few other resources. Now, researchers have sequenced the first complete Himalayan marmot genome, which may help them to better explain how the marmots live in such extremes.
The findings, which appear December 20 in the journal iScience, hint at the genetic mechanisms underlying high-altitude adaptation and hibernation, the researchers say. They also serve as a valuable resource for researchers studying marmot evolution, highland disease, and cold adaptation.
"As one of the highest-altitude-dwelling mammals, the Himalayan marmot is chronically exposed to cold temperature, hypoxia, and intense UV radiation," said Enqi Liu of Xi'an Jiaotong University Health Science Center in China. "They also hibernate for more than six months during the wintertime."
Those striking biological features led Liu and his team, including first author Liang Bai, to consider the Himalayan marmot as an ideal animal model for studying the molecular mechanisms of adaptation to extreme environments. To begin, they sequenced and assembled a complete draft genome of a male Himalayan marmot. They also re-sequenced 20 other Himalayan marmots, including individuals living at high and low altitudes, and four other marmot species. Additionally, RNA sequencing was done to compare gene-expression differences between marmots in a state of torpor and awake marmots.
The DNA data show that the Himalayan marmot diverged from the Mongolian marmot about 2 million years ago. The researchers identified two genes, Slc25a14 and ψAamp (a processed pseudogene), that have been selected in different directions in marmots living at low versus high altitudes, suggesting they are related to survival in high-altitude populations under conditions of extremely low oxygen.
They further suggest that Slc25a14 may have an important neuroprotective role. The shift in ψAamp affects the stability of RNA encoding the gene Aamp, which may be a protective strategy to prevent the excess growth of new blood vessels under extremely low-oxygen conditions.
The RNA sequencing data show that gene-expression changes occur in the liver and brain during hibernation. These include genes in the fatty acid metabolism pathway as well as blood clotting and stem cell differentiation.
Interestingly, a previous study suggested that because the hibernator's brain is exposed to near-freezing temperatures and has decreased blood flow, there is an increased risk of blood clots, the researchers note. Their brain stem cells may also be better prepared to repair injuries as an adaptation needed to survive extreme environmental stresses.
The researchers say they plan to continue improving the quality of the Himalayan marmot's genome. They note that the Himalayan marmot is also known for being highly susceptible to woodchuck hepatitis virus and is a natural host and transmitter of the plague to humans.
"We will elucidate the immune system features responsible for the hepatitis virus and bacterial infection," Liu said.
More information: iScience, Liu et al.: "Hypoxic and cold adaptation insights from the Himalayan marmot genome" www.cell.com/iscience/fulltext … 2589-0042(18)30224-4 , DOI: 10.1016/j.isci.2018.11.034
Provided by Cell Press