Deep learning democratizes nano-scale imaging
Many problems in physical and biological sciences as well as engineering rely on our ability to monitor objects or processes at nano-scale, and fluorescence microscopy has been used for decades as one of our most useful information sources, leading to various discoveries about the inner workings of nano-scale processes, for example at the sub-cellular level. Imaging of such nano-scale objects often requires rather expensive and delicate instrumentation, also known as nanoscopy tools, which can only be accessed by professionals in well-resourced labs.
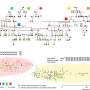
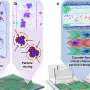
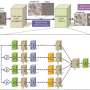
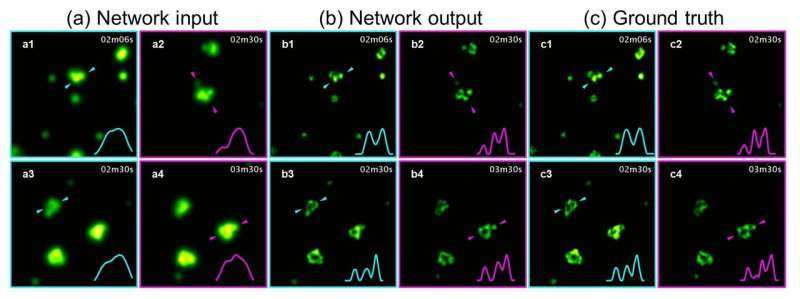
To democratize access to high-resolution fluorescence imaging and be able to resolve and monitor objects at nano-scale, UCLA researchers have developed a new method, based on artificial intelligence, to digitally transform fluorescence images acquired using a lower resolution and simpler microscope into images that match the resolution and quality of higher resolution and advanced microscopes that are built for nano-scale imaging. To achieve this transformation, an artificial neural network is trained by thousands of image pairs (lower resolution vs. higher resolution images of the same samples), teaching the deep neural network the cross-modality image transformation from a much simpler and cheaper microscope into a high-end nanoscope. Once the training is complete, the deep neural network can blindly take in an image of the lower resolution and simpler microscope to digitally super-resolve the features of the nanoscopic objects in the sample, matching the performance of a much more advanced nanoscopy instrument.
This work was published in Nature Methods, a journal of the Springer Nature Publishing Group. This research was led by Dr. Aydogan Ozcan, an associate director of the UCLA California NanoSystems Institute (CNSI) and the Chancellor's Professor of electrical and computer engineering at the UCLA Henry Samueli School of Engineering and Applied Science. Hongda Wang, a UCLA graduate student, and Yair Rivenson, a UCLA postdoctoral scholar, are the study's co-first authors.
This nanoscopic image transformation framework builds bridges across different imaging modalities and instruments, and its success was demonstrated by super-resolving various biological cells and tissue samples, matching the imaging resolution of much more advanced fluorescence nanoscopy tools using much simpler and more accessible microscopes. Furthermore, this technique allows imaging of dynamic events at nanoscale over a much larger sample volume, while also reducing the toxic effects of illumination photons on living organisms and cells.
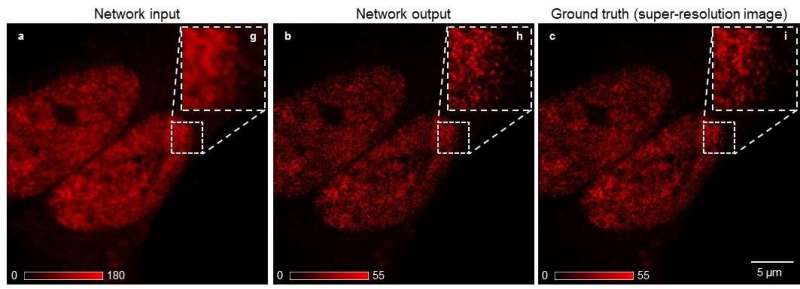
"Our work demonstrates a significant step forward in computational microscopy, which might help to democratize super-resolution imaging by enabling new biological observations at nano-scale beyond well-equipped laboratories and institutions," said Ozcan.
Other members of the research team were Yiyin Jin, Zhensong Wei, Ronald Gao, Harun Günaydin, members of the Ozcan Research Lab at UCLA, as well as Dr. Laurent A. Bentolila, the Director of CNSI Advanced Microscopy Facility at UCLA and Dr. Comert Kural, an assistant professor at the department of physics at the Ohio State University.
Ozcan lab is supported by NSF, HHMI and Koc Group. Imaging experiments were performed at the Advanced Light Microscopy/Spectroscopy Laboratory at CNSI and at the Advanced Imaging Center at Janelia Research Campus.
More information: Hongda Wang et al. Deep learning enables cross-modality super-resolution in fluorescence microscopy, Nature Methods (2018). DOI: 10.1038/s41592-018-0239-0
Journal information: Nature Methods
Provided by UCLA Ozcan Research Group