Mechanism safeguarding unique epigenome of oocytes and maternal fertility

In mammals, females have a limited supply of oocytes. Oocytes have a unique epigenome with approximately half the DNA methylation of sperm, and the most terminally differentiated somatic cells. Until recently, regulators of this unique DNA methylation pattern and its functional significance were unknown.
Now, a novel DNA methylation regulator called Stella has been identified, whose ectopic overexpression in somatic cells leads to global DNA demethylation by disrupting the function of the DNA methylation regulator UHRF1.
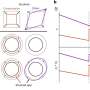
In a recent study published online ahead of print in Nature, a joint research group led by Dr. Zhu Bing from the Institute of Biophysics of the Chinese Academy of Sciences reveals that Stella sequesters UHRF1 from the nucleus through an active nuclear export process, and the dysregulation of UHRF1 by loss of Stella results in an accumulation of aberrant DNA methylation during postnatal oogenesis. These findings show the first regulatory factor found to safeguard the unique methylation status of the oocyte genome.
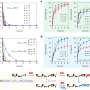
Since Stella is highly expressed in oocytes, the researchers focused on the in vivo function of Stella during oogenesis. Earlier studies revealed that Stella-null oocytes were incapable of supporting the development of preimplantation embryos. This study shows preferential hypermethylation at the transcriptionally inert regions of Stella null oocytes. These aberrant promoters of hypermethylation on the maternal allele severely affected zygotic genome activation and development of the preimplantation embryo.
Interestingly, a maternal genome lacking DNA methylation had been reported not to affect preimplantation embryo development, while this study suggests that keeping a uniquely hypomethylated oocyte genome is vital.
Moreover, researchers found that DNMT1, generally considered to be a maintenance DNA methyltransferase, which is only active on hemi-methylated DNA in vivo, is the major DNA methyltransferase responsible for the aberrant DNA methylation in Stella-deficient oocytes and unambiguously proves the de novo methylation activity of DNMT1 in vivo.
This discovery rewrites the textbook classification of DNA methyltransferases. Also, it sheds light on a functional role of DNMT1 in post-mitotic cells, which may help to reveal a role for DNMT1 in ageing.
More information: Yingfeng Li et al, Stella safeguards the oocyte methylome by preventing de novo methylation mediated by DNMT1, Nature (2018). DOI: 10.1038/s41586-018-0751-5
Journal information: Nature
Provided by Chinese Academy of Sciences