Tweaking just a few genes in wild plants can create new food crops – but let's get the regulation right

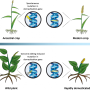
The crops we rely on today have been bred over thousands of years to enhance certain characteristics. For example, sweetcorn started life as a wild grass called teosinte.
But every time we select for a trait through breeding – such as repeatedly crossing selected plants to produce bigger fruits – we lose genetic diversity which is the essential variation for other traits like disease resistance. This leaves our crops vulnerable to pests and disease.
Precise gene editing technologies could offer a solution.
CRISPR gene editing has been successfully used to re-domesticate wild tomato plants. One research group edited only six genes and produced a commercially sized fruit in a wild relative of tomato, Solanum pimpinellifolium. Another group achieved a similar result by editing only four genes.
As a society we need to work out how such technology and plants will be regulated to ensure safety and acceptability.
The new domestication
The new approach, termed "de novo domestication" or "new domestication", allows the genetic diversity of the wild plant to feature in a new crop. These new tomato lines retain all of the diversity of their ancestors, providing protection against disease.
They taste good too, apparently.
This was possible thanks to years of painstaking research into the genes that underpin the essential traits related to domestication. Without this, scientists wouldn't know which genes to target and edit.
Some of the genes were identified by crossing plants with different traits (like large versus small fruits). Others were discovered by comparing wild relatives with domesticated plants.
Could we do this with other wild species?
We currently rely on very few plant species for the majority of the world's food production. More than half of our plant-derived energy intake comes from just three grasses (wheat, rice and corn). Gene editing could provide a way to expand this.
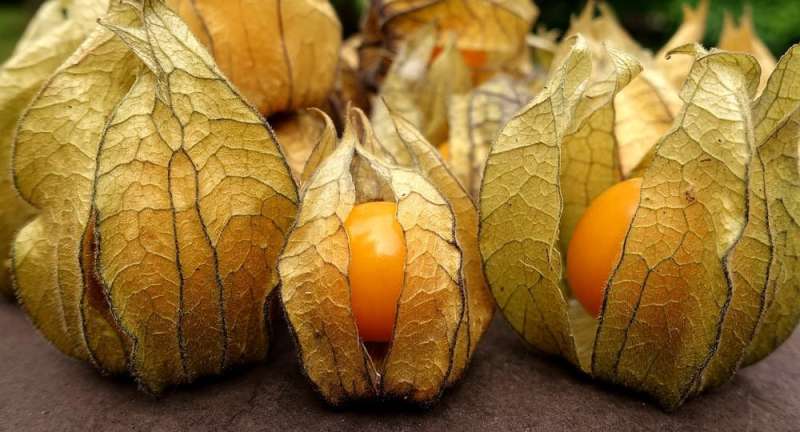
Showing the broader value of the tomato approach described above, another research group applied the same method to an orphan crop Physalis pruinosa (known as the ground cherry). Orphan crops are those that have been neglected and escaped modern agriculture for various reasons. They receive little investment, research or breeding effort.
The researchers hope the ground cherry will one day find its place alongside the strawberry, blueberry, blackberry and raspberry in large-scale agriculture.
The "de novo domestication" approach potentially provides a way to domesticate any edible wild plant. With an estimated 20,000 known edible species, the possibilities for domestication could be extraordinary – particularly in Australia, where broad, economically successful crop domestication of native foods has been mostly limited to the macadamia nut.
Gene sequencing getting cheaper
To create new crops, we need good working knowledge of the gene targets, and the genome sequence (which contains the complete code of all genes inside each cell) of the plant species that we want to domesticate.

Genome sequencing used to cost many millions of dollars and require massive research teams. It's now increasingly cheap and routine.
Our understanding of the target genes comes largely from studies of major agricultural crops. Some domestication genes will only work in species that are closely related to the crop in which they were discovered.
Versions of the domestication genes targeted in the tomato studies are found in many plant species, so the approach might well work in more distantly related species too.
How should we regulate this?
We should be fostering this kind of innovation, but we need to do it safely.
Worldwide, policymakers have wrestled with the implications of the new genetic tools. Debate has sprung up around how to regulate genome editing, compared with existing genetic modification methods.
Editing genes in a crop is different to traditional genetic modification, or transgenics – in which a gene from a different species is inserted into a plant.
In contrast to both of these approaches, classic crop breeding has relied on random processes like irradiation to induce new genetic diversity. CRISPR editing is similar but more efficient and precise, because it targets a specific desired mutation.
In July, European courts ruled that edited plants fall under the same regulation as transgenics. This places them under very strict regulations that create significant hurdles to enter the market, potentially driving talent and funding out of Europe.
In contrast, the US Department of Agriculture (USDA) said it would not regulate genome-edited crops.
A balanced approach
A reasonable balance between these two regulatory approaches is probably the most sensible way forward. Genome editing shouldn't completely escape regulation.
If it can be demonstrated that the edited plant doesn't contain any new genes (including CRISPR machinery) then the regulation should be much less stringent than for transgenics, because the changes are so similar to conventional plant breeding. Sequencing the genome of the edited crop is a good way to provide evidence of this.
In Australia, genetically modified organisms are regulated by the Office of the Gene Technology Regulator (OGTR). The current legislation defines genetic modification very broadly, but is under review. South Australia is an exception to this, with a ban on genetically modified crops.
Ideally, regulation should focus more on questions around the types of genetic modifications that we should allow in our crops than the way that they were introduced and where they came from.
But edited organisms shouldn't be completely excluded from regulation. Evidence should be requested, and provided, that new crops are functionally equivalent to the products of conventional breeding and the subsequent approval process should reflect this.
The primary priority for policymakers and regulators is to ensure crop safety. Maintaining an open and transparent dialogue will be crucial so that the public can trust the decisions.
Provided by The Conversation
This article is republished from The Conversation under a Creative Commons license. Read the original article.![]()