The hidden hazards of antibiotic resistance genes in air

People are often notified about poor air quality by weather apps, and this happens frequently in urban areas, where levels of outdoor pollution containing particulates and soot are high. But now scientists are reporting in ACS' Environmental Science & Technology that there is another type of air contaminant that they say isn't receiving enough attention: antibiotic-resistance genes.
According to the U.S. Centers for Disease Control and Prevention, at least 2 million people in the U.S. become infected with antibiotic-resistant bacteria every year. Research has shown that antibiotic resistant genes (ARGs) can move from bacteria to bacteria, or even from bacteria to the environment. For example, tetracycline-resistance genes have been found near animal feed operations, and β-lactam-resistance genes have been found in urban parks in California. These studies indicated that airborne transmission could be a factor in the spreading and exposure of ARGs. But current air pollution investigations typically don't take ARGs into account. So, Maosheng Yao and colleagues wanted to examine airborne ARGs on a global scale.
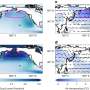
The team performed a survey of 30 ARGs across 19 cities around the world, including San Francisco, Beijing and Paris. The group studied ARGs resistant to seven common classes of antibiotics: quinolones, β-lactams, macrolides, tetracyclines, sulfonamides, aminoglycosides and vancomycins. Beijing had the most diverse group of airborne ARGs, with 18 different subtypes detected, while San Francisco had the highest overall level of airborne ARGs. Genes resistant to β-lactams and quinolones were the two most abundant types of ARGs in all the cities studied. Low levels of ARGs resistant to vancomycin, an antibiotic of last resort for MRSA treatment, were found in the air of six cities.
More information: Global Survey of Antibiotic Resistance Genes in AirEnvironmental Science & Technology (2018). pubs.acs.org/doi/abs/10.1021/acs.est.8b02204
Abstract
Despite its emerging significant public health concern, the presence of antibiotic resistance genes (ARGs) in urban air has not received significant attention. Here, we profiled relative abundances (as a fraction, normalized by 16S rRNA gene) of 30 ARG subtypes resistant to seven common classes of antibiotics, which are quinolones, β-lactams, macrolides, tetracyclines, sulfonamides, aminoglycosides, and vancomycins, in ambient total particulate matter (PM) using a novel protocol across 19 world cities. In addition, their longitudinal changes in PM2.5 samples in Xi'an, China as an example were also studied. Geographically, the ARGs were detected to vary by nearly 100-fold in their abundances, for example, from 0.07 (Bandung, Indonesia) to 5.6 (San Francisco, USA). The β-lactam resistance gene blaTEM was found to be most abundant, seconded by quinolone resistance gene qepA; and their corresponding relative abundances have increased by 178% and 26%, respectively, from 2004 to 2014 in Xi'an. Independent of cities, gene network analysis indicates that airborne ARGs were differentially contributed by bacterial taxa. Results here reveal that urban air is being polluted by ARGs, and different cities are challenged with varying health risks associated with airborne ARG exposure. This work highlights the threat of urban airborne transmission of ARGs and the need of redefining our current air quality standards in terms with public health.
Journal information: Environmental Science & Technology
Provided by American Chemical Society