How the flu virus builds a better mousetrap
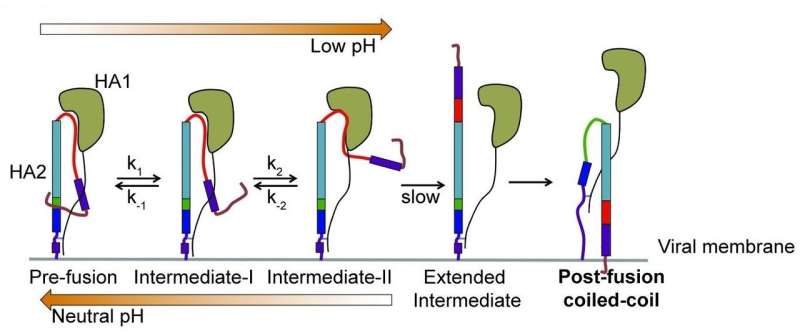
For the first time, scientists have directly visualized in real-time structural changes in the surface protein of the influenza virus that may help the virus to fuse with and enter target cells before hijacking their functions. Researchers at Tufts University School of Medicine found that single molecules of the protein hemagglutinin (HA) that reside on the surface of the virus unfold to stretch toward target cells, then refold and try again 5 to 10 times per second. The discovery shows the flu virus to be more dynamic than previously thought and may help efforts to develop more effective vaccines and better understand other viruses such as Ebola, HIV, and SARS. The research appears in the journal Cell online June 28 and in print August 9.
For decades, influenza has served as the study model for a large class of viruses that enter cells by a common mechanism: An envelope protein on the surface of these viruses must attach the virus to the cell membrane, and then fuse the virus and the cell. Fusion allows release of the virus contents into the cell, so it can take over the cell's internal functions and replicate. Influenza's envelope protein, HA, has long been a template for fusion mechanisms in other viruses.
"Envelope proteins have been described as old-fashioned mousetraps, set in a static, spring-loaded state, waiting to be triggered by interaction with a target cell," said the study's senior author, James Munro, Ph.D., assistant professor of molecular biology and microbiology at Tufts School of Medicine who also teaches at the Sackler School of Graduate Biomedical Sciences at Tufts. "Once triggered, they undergo a dramatic change in their three-dimensional structure, enabling fusion and entry into the target. However, despite some hints in previous research, this process hadn't been directly observed, and it was widely thought that each protein molecule on the surface of the virus had only one chance to spring its trap."
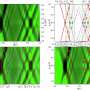
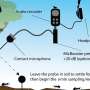
Using an advanced imaging technology—single-molecule Förster resonance energy transfer, or smFRET, which measures nanoscale distances within single molecules labeled with fluorescent dyes—and then performing significant computational analyses of the data, the Tufts researchers generated the first real-time visualization of the changing shape of individual HA molecules seeking cellular targets. To facilitate the experiments, the HA molecules were imaged while on the surface of an unrelated virus.
What they discovered was a versatile and dynamic mousetrap that was far from the one and done model previously assumed. "The fact that this viral molecule can reconfigure itself, then reverse that configuration and rapidly repeat that sequence multiple times changes the way we think about virus entry," said Munro.
Reversibility may potentially benefit the virus in several ways, including preventing early activation in the absence of an appropriate target, enabling virus molecules to synchronize their efforts to increase efficiency, and confusing a cell's protective antibodies, which must recognize the shape of a virus in order to defend against it.
"Surface proteins are the only part of the virus that the immune system 'sees.' As a result, nearly all known antibodies that inhibit virus replication target these proteins," noted Munro. "We are asking 'What structures does the immune system need to recognize to make more effective antibodies?'."
Research is still needed to prove repeatability and reversibility of these protein dynamics in viruses other than flu, and visualization experiments using inert, non-infectious Ebola particles are underway in Munro's lab. Munro is the recipient of a National Institutes of Health Director's New Innovator Award to support use of single-molecule imaging to investigate how viruses such as Ebola enter host cells.
More information: Das, D., Govindan, R., Nikic-Spiegel, I., Krammer, F., Lemke, E., Munro, J. (2018) Direct visualization of the conformational dynamics of single influenza hemagglutinin trimers. Cell. DOI: 10.1016/j.cell.2018.05.050
Journal information: Cell
Provided by Tufts University