New nuclear RNA retention activity discovered
Gene expression involves mRNA transport from its place of synthesis to the cytoplasm where protein translation occurs. However, many non-coding RNA species do not follow this flow and new data now demonstrate how cells prevent the unwanted export of RNA and instead ensure nuclear degradation.
Mammalian genomes produce a large repertoire of RNAs, which experience very different fates. Protein-coding mRNAs serve as a prototypic example of transcripts that are rapidly processed and exported from nucleus to the cytoplasm to translate protein. However, several RNAs, many of which are non-coding, are rapidly degraded in the nucleus by the highly conserved RNA degradation machine, the RNA exosome.
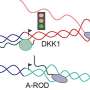
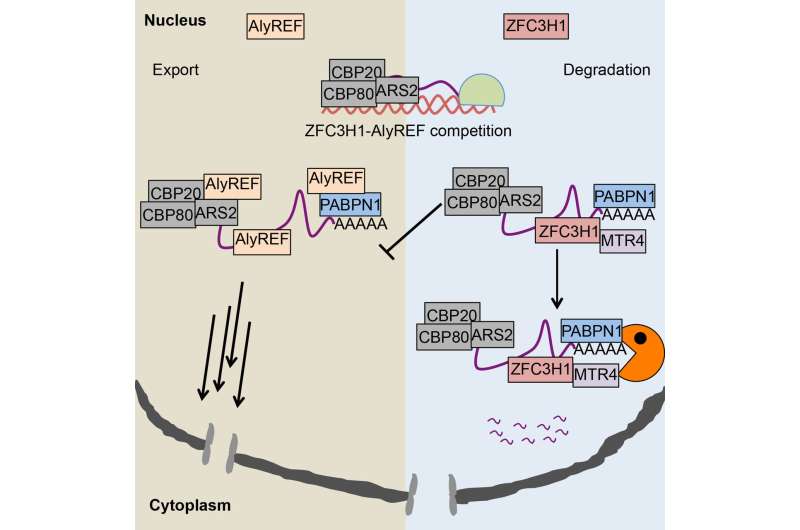
In recent work, postdoctoral researcher Toomas Silla from Torben Heick Jensen's laboratory at the Department of Molecular Biology and Genetics discovered that abolished RNA exosome function causes the accumulation of polyadenylated (pA+) RNAs in distinct nuclear foci (coined as pA+ foci), indicating that these accumulated RNAs are prevented from entering the cytoplasm. This hinted that so-called exosome adaptors, which are required for RNA exosome loading onto its substrates, might dually serve as RNA nuclear retention factors and counteract export activity.
Previously, the Torben Heick Jensen laboratory established the Nuclear EXosome Targeting (NEXT) complex and PolyA eXosome Targeting (PAXT) connection to be adaptor complexes that link the RNA exosome to short unprocessed or to longer polyadenylated RNAs, respectively (Lubas et al. Mol Cell 2011; Meola et al. Mol Cell 2016). Interestingly, co-localization analysis revealed, that pA+ foci are particularly enriched for PAXT proteins such as MTR4, ZFC3H1 and PABPN1. Among these factors, ZFC3H1 appeared to be critical for establishing the pA+ RNA foci; in ZFC3H1 depleted cells pA+ RNA foci are not formed, and exosome targets that would normally be degraded in the nucleus, are exported to the cytoplasm. These transcripts now enter into the RNA export pathway, demonstrating that ZFC3H1 counteracts RNA export activity to ensure the nuclear retention of exosome targets before their decay.
Transcriptome-wide characterization of pA+ RNA foci contents, revealed surprisingly many full-length and spliced mRNAs. Thus, in addition to bona fide RNA exosome targets, mRNAs may also be subjects to nuclear decay. Although most mRNAs are quickly exported after their production, these new results therefore suggest that mRNAs can be cut out of the export pathway and degraded in the nucleus instead. Future research will be focused on determining how sorting between productive export and retention/decay is orchestrated and which additional factors contribute to this process.
The scientific article was published in the internationally recognised journal Cell Reports.
More information: Toomas Silla et al. The RNA Exosome Adaptor ZFC3H1 Functionally Competes with Nuclear Export Activity to Retain Target Transcripts, Cell Reports (2018). DOI: 10.1016/j.celrep.2018.04.061
Journal information: Cell Reports
Provided by Aarhus University