Genome Jenga study reveals unexpected gene alliances in the cell
To understand how a cell works, biologists like to take it apart. By removing genes from cells in diverse combinations, researchers have now uncovered how different genes work together to keep cells alive. The research will help scientists understand how faults in multiple genes combine to drive common diseases such as cancer or heart disease.
Led by Charles Boone, a professor in the University of Toronto's Donnelly Centre, Brenda Andrews, University Professor and director of the Donnelly Centre, and Professor Chad Myers, of the University of Minnesota-Twin Cities, MN, the research builds up on the teams' previous work which showed how genes combine in pairs to underpin cell's health. Taking it a step further, the new study examines for the first time how higher-order gene combinations-comprising three genes- help maintain normal cell physiology, as revealed in Science.
Boone and Andrews are also professors in U of T's Department of Molecular Genetics and Senior Fellows at the Canadian Institute for Advanced Research (CIFAR) and Myers is a Fellow at CIFAR.
"There's a growing understanding that interactions between genes can drive inherited disease susceptibility, which is why we have to understand the general principles of these genetic interactions," says Boone.
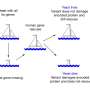
It's very much like a giant game of Jenga, with thousands of gene blocks that can be removed. While most single blocks can be taken out without compromising the structure, when critical combinations of blocks are removed, the system collapses. Similarly, genes with different roles can combine to keep the cell alive. By unpicking such gene alliances, scientists hope to reveal clues about the foundations of personal health.
It's now clear from genome sequencing studies that each person carries thousands of genetic variants—differences in genes' DNA sequence—that could combine to impact our health. However, these studies do not have the statistical power to predict a person's risk of disease from their unique combination of genetic variants. This poses a major obstacle for personalized medicine which seeks to use genome information to predict risk of disease and tailor treatment.
To uncover the rules of combinatorial gene function, the team previously investigated how genes work in pairs in yeast cells. The yeast is one of biologists' favourite cell models due to its relatively small genome comprising 6,000 genes and an already existing wealth of data. Having previously removed from yeast all possible gene pairs-18 million of them- the team now went a step further to examine what happens when you remove a subset of 36 billion possible trigenic combinations.
They found that, similar to interactions between two genes, trigenic interactions also mainly occur between genes that are functionally related- they code for parts belonging to the same molecular machine or that exist in the same part of the cell, for example. But with trigenic interactions, the researchers also began to see more surprising partnerships between genes that have unrelated function and are involved in different bioprocesses in the cell.
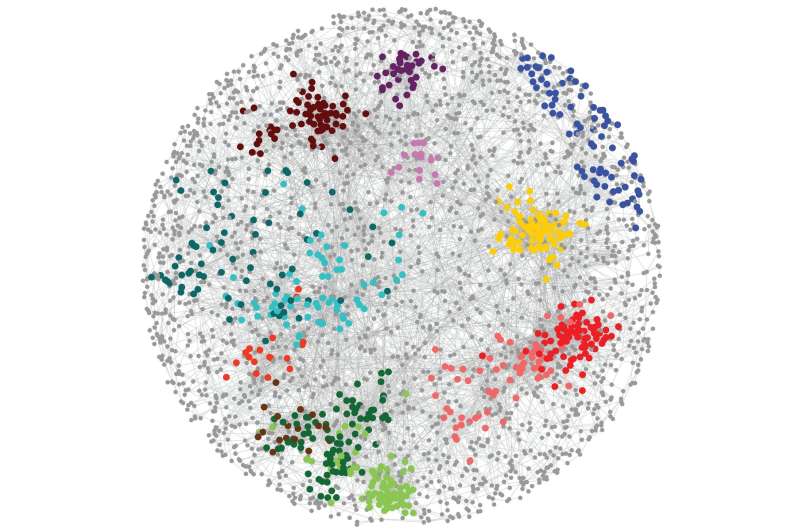
"Studying genetic networks allows you to see how genes are connected, how biological processes talk to one another and how a cell deals with perturbations in multiple genes," says Elena Kuzmin, a lead author on the paper and a previous graduate student in the Boone lab who is now a postdoctoral fellow at McGill University in Montreal. "You get a global view of the cell," she says.
Furthermore, using mathematical modeling the researchers estimate that all genes in the cell have a role to play when trigenic interactions are taken into account. This could finally explain why only a tenth of yeast's 6,000 genes are essential for cell survival, a rule that holds for other cell types including human cells.
Thanks to recent advances in gene editing, it is now possible to remove combinations of genes from human cells, which Boone and Andrews labs are currently doing in collaboration with Jason Moffat's group in the Donnelly Centre to map relationships between disease genes.
"Our yeast work demonstrates how mutations in multiple genes combine to have unexpected effects and is providing a roadmap for understanding genetic interactions in much more complex cells and organisms, including humans." says Andrews. "Identifying combinations of genes that work together to underpin robust biological systems is important for deciphering what goes wrong with its collapse into a disease state."
More information: "Systematic analysis of complex genetic interactions" Science (2018). science.sciencemag.org/cgi/doi … 1126/science.aao1729
Journal information: Science
Provided by University of Toronto