Reading the entire human genome – one long sentence at a time

Fifteen years ago, the Human Genome Project announced they had cracked the code of life. Nonetheless, the published human genome map was incomplete and parts of our DNA remained to be deciphered. Now, a new study published in the journal Nature Biotechnology brings us closer to a complete genetic blueprint by using a nanotechnology-based sequencing technique.
Like ancient Egyptian ruins covered in mysterious hieroglyphics, the letters and words in our genetic code remained unutterable for a long time. In an effort to solve this genetic cipher, the Human Genome Project, a collaborative international consortium, was created. The goal was to read out the DNA sequence – made up of four letters, or bases, A,T,G and C – of all human genes (genome). In 2003, a near-complete map of the human genome was reported. The scientific community hailed the momentous event as a turning point, perhaps overshadowed only by the discovery of the double-helix structure of DNA. Indeed, for the first time in human history, we could read and understand the language of our "being". Yet, the assembled genome represented only 92% of all human genes. Gaps remained that could not be easily decrypted. For many researchers, that elusive 8% of the genome is a holy grail.
The dark matter inside us all
The unmappable genome is associated with "heterochromatin" (dark matter of the genome, highly condensed), unlike "euchromatin" (light matter, more loosely wound part of the genome). Euchromatin is gene-rich while heterochromatin refers to the silent, repressed regions of our DNA. Euchromatin is full of unique DNA sequences. This means that finding a single- or low-copy DNA sequence, with all the same DNA bases in the same order, at more than one location in our genome is highly unlikely. These discrete DNA sequences are easily distinguishable and serve distinct purposes within our cells. No wonder the human genome has almost 20,000 different genes with limited redundancy. Now, visualize a human chromosome as a big "X", made of coiled-up DNA, with two arms attached at a constriction. Heterochromatin is mostly localised near the point of attachment (centromere) and the tips of the arms (telomeres). In fact, the centromere becomes indispensable when cells divide, dragging along one chromosome arm into each of the newly formed daughter cells.
DNA sequencing technologies operate by reading each base of DNA, one at a time, and spitting out short "reads" that spell out the sequence being read. Thus, decoding unique, non-identical euchromatic DNA is facile because one stretch apart from other with little ambiguity. The problem arises when we try to enunciate heterochromatic sequences comprising strings of DNA that look like each other. Arranged in tandem arrays or dispersed throughout our genome, these highly repetitive stretches of DNA amount to garbled gibberish after conventional DNA sequencing. One small chunk of DNA (monomer) at the centromere resembles other identical chunks flanking it and so on. In the resulting quagmire, the base-composition & precise position of any given repeated sequence cannot be ascertained in a long polymer of repeats. Made up of millions of repeating A,T,G,C bases, the centromeres of human chromosomes evaded biologists and explain holes in our current DNA map.
Threading the genome into a tiny needle
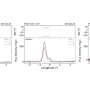
The new study, from the team of Dr. Karen Miga at University of California (Santa Cruz), has managed to uncover the centromere of the Y chromosome – the male-specific chromosome and also the smallest chromosome in our genome (something worth thinking about). The researchers were able to insert a longer stretch of DNA into a nano-pore (like thread passed through the eye of a needle), "resulting in complete, end-to-end sequence coverage of the entire insert". Using this nanopore-sequencing method, the researchers can now decipher a long, muddled DNA stretch full of repeats. This "long-read" strategy allowed them to string together longer pieces of DNA (made up of variable repeat monomer lengths). It turns out that when all these chunks are laid out, certain clues help reconstruct the repetitive-sequence. Walking along the centromere, from left to right, context is provided by surrounding monomers in the same tandem array and by flanking non-repetitive DNA.
Like a neatly laid section of railroad, the authors pieced together a chain of contiguous DNA sequences and solved the jigsaw puzzle of the Y chromosome centromere. This recent work, published in Nature Biotechnology journal, plugs holes in the existing human DNA map. In the future, finding out the DNA sequences that define other centromeres will allow researchers to rewrite, manipulate, alter or duplicate these key structures. Given that the centromere is essential for cells to divide and segregate their genetic content to future generations, the Y centromere assembly represents an exciting step forward in modern biology.
More information: Miten Jain et al. Linear assembly of a human centromere on the Y chromosome, Nature Biotechnology (2018). DOI: 10.1038/nbt.4109
Journal information: Nature Biotechnology
Provided by The Conversation
This article was originally published on The Conversation. Read the original article.![]()