Designing microbial communities to help plants battle nutritional stress

Plants and microbes engage in a diverse array of symbiotic relationships, but identifying the specific microbes or groups of microbes that contribute to plant health is extremely difficult. In work published on February 20th in the open access journal PLOS Biology, researchers devised a general experimental scheme to identify and predict which small groups of bacterial species can help plants respond to phosphate starvation, a form of nutritional stress.
Microbes can provide an astonishing range of beneficial functions to plants through their colonization of roots and above-ground organs such as leaves. Collectively, these microbes comprise the so-called "plant microbiome." The hope of several research groups and biotech companies around the world is to define and deploy single microbes, or small groups of microbe species, that can help plant productivity and lessen the reliance on chemical fertilizers in agriculture.
While researchers can easily define "who's there" (i.e. what kinds of bacteria and fungi are present) and in what relative proportions, it is more difficult to know how those microbes interact with each other and with the plant to influence plant performance. Jeff Dangl, co-corresponding author of the article, hopes that their work will "provide a template for researchers interested in defining probiotics with predictive effects on both plants and humans."
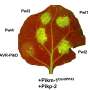
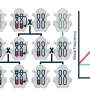
In this research article, co-first authors Sur Herrera Paredes and Tianxiang Gao, co-corresponding author Gabriel Castrillo and their colleagues from the Howard Hughes Medical Institute and the departments of biology and computer science at the University of North Carolina at Chapel Hill, devised a general scheme to build a small consortium of eight or nine bacterial members based on results from laboratory experiments. They first tested single strains of bacteria to determine which bacteria helped or hindered the plant's ability to survive in phosphate starvation. They then tested a small subset of all the possible pairs of eight or nine-membered consortia made from these single strains, used those results to predict which untested pairs of consortia would help the plant using a Neural Network analysis, and validated the predictions experimentally.
This research demonstrates that the functional complexity of the root microbiome might be simplified by testing only a subset of all possible microbial combinations.
"The hope of future research is that it should be possible to build plant probiotics with predictable impacts on our plants, and similarly, on our own human microbiome," Dangl said.
More information: Herrera Paredes S, Gao T, Law TF, Finkel OM, Mucyn T, Teixeira PJPL, et al. (2018) Design of synthetic bacterial communities for predictable plant phenotypes. PLoS Biol 16(2): e2003962. doi.org/10.1371/journal.pbio.2003962
Journal information: PLoS Biology
Provided by Public Library of Science