Promise of new antibiotics lies with shackling tiny toxic tetherballs to bacteria
Biologists at The University of Texas at Austin have developed a method for rapidly screening hundreds of thousands of potential drugs for fighting infections, an innovation that holds promise for combating the growing scourge of antibiotic-resistant bacteria. The method involves engineering bacteria to produce and test molecules that are potentially toxic to themselves.
A description of the method appears in the Jan. 25 print edition of the journal Cell.
No new class of antibiotic has been discovered in 40 years—many of the most accessible ones in nature already have been found, and the process for creating and testing new ones from scratch is slow and laborious—but modern medicine is in sore need of them. According to the World Health Organization, antibiotics have added about 20 years to the average human lifespan. But their protective benefits are slipping away as bacteria evolve antibiotic resistance.
In their proof of concept, the UT Austin team, led by Bryan Davies, screened about 800,000 molecules called peptides to see whether they had antimicrobial effects, meaning they killed harmful bacteria. Of those, several thousand killed E. coli bacteria, making them potential leads for antibiotics. Some antibiotics currently in use are peptides. Follow-up research will be necessary to determine which, if any, of the thousands of new leads are truly effective and safe in humans, but the researchers demonstrated that at least one such molecule, dubbed P7, also kills other forms of pathogenic bacteria and is safe in mice.
With this method, called SLAY (Surface Localized Antimicrobial Display), one person can screen hundreds of thousands of similar peptides faster and more cost-effectively than existing methods can. Davies would like to see the method become a standard tool in the global hunt for new antibiotics.
"So what if we have a thousand groups all using this system to follow their own interests and their own peptides?" said Davies, assistant professor of molecular biosciences. "Once you enable a community of that size, then I think you have a better chance of actually finding a new antibiotic that works."
A key advance in this work was figuring out how to get bacteria to produce molecules that might be toxic to themselves and to control how those molecules interact with their host bacteria.
"We thought, wouldn't it be great if a bacteria could synthesize the compound for us, because bacteria are cheap and easy to grow, and then test the compound on itself and report back and tell us, was that an antimicrobial or not?" Davies said.
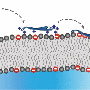
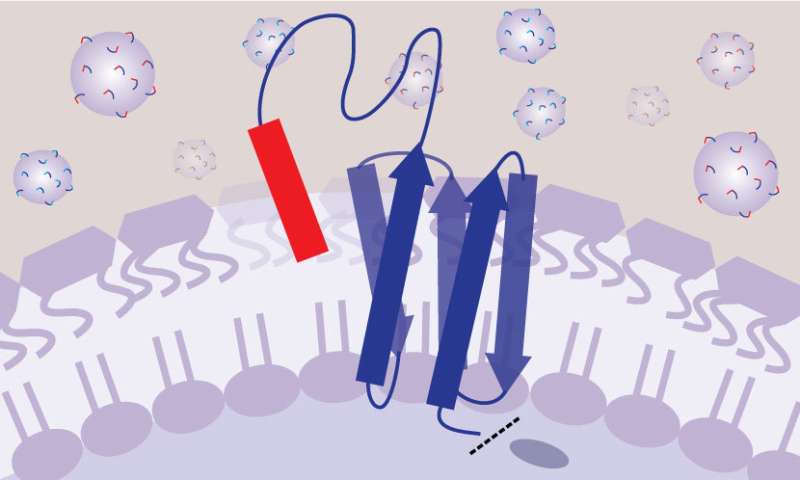
Their solution was to genetically engineer the bacteria to produce a molecule on the cell surface that is part peptide and part tether—like a playground tetherball and its tether—with one end fixed to the cell membrane and the other end free to float around. This allows the peptide to move around and make contact with the bacterial cell surface, as if it were free-floating like a drug in your bloodstream, but without interacting with other nearby bacteria.
By ensuring that each version of the tetherball only interacts with the bacteria that produced it, the researchers could then make a big leap in efficiency. They could create hundreds of thousands of strains of bacteria—each genetically engineered to produce a slightly different version of the tetherball—and put all of these strains into the same test tube to grow. By running hundreds of thousands of experiments simultaneously, their method saves a tremendous amount of space, time and cost.
Part of this process relies on a technique developed by UT Austin's George Georgiou in the 1990s that induces bacteria to produce proteins or peptides on their surfaces.
To find out which tetherballs (peptides) knock out their hosts, the scientists use gene sequencing to identify which versions are being produced by bacteria at the start and which are being produced at the end.
Following on the discovery that P7 kills pathogens, the team now plans to create thousands of subtle variations of this molecule, called derivatives, and run them through the same screening process to search for an even more effective version.
Postdoctoral fellow Ashley Tucker led the experimental work to demonstrate use of the platform.
Davies, Tucker and UT Austin have filed patent applications for the SLAY method and for the specific genetic sequences for the thousands of antimicrobial peptides they have discovered so far.
More information: Ashley T. Tucker et al, Discovery of Next-Generation Antimicrobials through Bacterial Self-Screening of Surface-Displayed Peptide Libraries, Cell (2018). DOI: 10.1016/j.cell.2017.12.009
Journal information: Cell
Provided by University of Texas at Austin