The toxic sugar tree: Mapping the evolutionary history of a cancerous sugar gene

Around two million years ago, a genetic change occurred that differentiated humans from most other primates that both protected humans from diseases, yet made red meat a health risk.
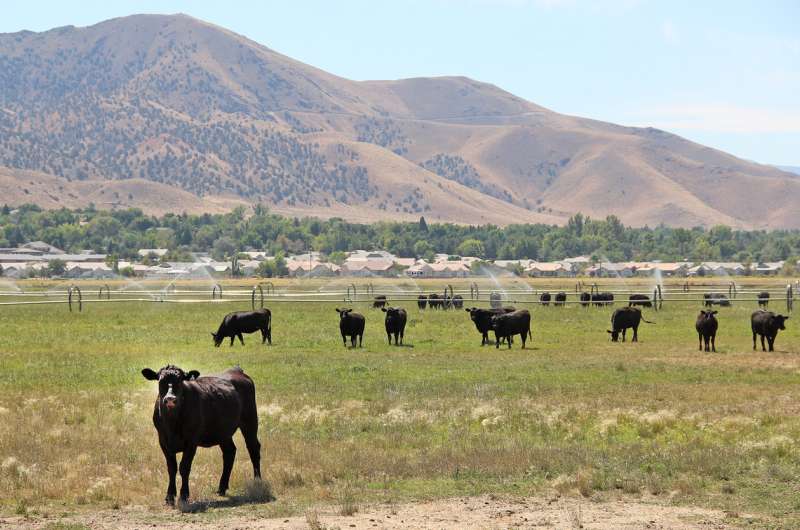
At this point in human evolution, a certain gene, known as CMAH, that allows for the synthesis of a sugar called Neu5Gc, went missing. This sugar is present in red meats, some fish and dairy products. When humans consume an animal that has that gene, the body has an immune reaction to the foreign sugar, which can cause inflammation, arthritis, and cancer.
University of Nevada, Reno researchers, led by College of Science Assistant Professor David Alvarez-Ponce, have analyzed 322 animal genome sequences from the National Center for Biotechnology Information looking for animals that show the presence of active CMAH genes. They placed the data from the 322 animal genomes into a "tree" to determine when in an animal's evolutionary history did the CMAH gene became inactive or "turned off." This is useful in explaining why certain species have an active CMAH gene and why similar species don't.
The Alvarez-Ponce lab specializes in studying the evolution of genes and genomes using bioinformatics. When it comes to the few fish investigated so far, there is an insignificant concentration of the Neu5Gc sugar to be a medical concern, but the concentrations of Neu5Gc are significantly higher in fish eggs, better known as caviar. As masters' student Sateesh Peri puts it, "one of the most expensive foods is among the ones with the highest amount of toxic Neu5Gc."
Birds also lack active CMAH genes, implying that consuming animals like chicken, turkey, and geese is not associated with the harmful side effects. Another class of animals that do not have the CMAH gene are reptiles, except for one species of lizard. The presence of the gene in this lizard challenges prior propositions that the gene may have been lost in an ancestor of reptiles and birds.
Aside from dietary issues, the CMAH gene also proves to be a major factor in whether or not a transplanted organ from an animal would be accepted by a human because of the gene. When an animal with a CMAH gene's organs are transplanted into a human who does not have the CMAH gene, the human body might reject the organ because of the presence of the Neu5Gc sugar.
"Inactivation of CMAH during human evolution might have freed humans from a number of pathogens," Alvarez-Ponce said. "For instance, there's this certain type of malaria that requires Neu5Gc for infection - other primates are susceptible to it, but not humans."
The presence of the CMAH gene determines what animals humans should avoid eating (or should eat in moderation) and which animal bites humans should avoid. If the animal has the CMAH gene, then it shouldn't be eaten because it could cause inflammation, arthritis and cancer. However, consuming red meat in moderation is considered to be fine. If the animal doesn't have the CMAH gene, then those animals are more likely to contain pathogens that attach to Neu5Ac (the precursor of Neu5Gc) and that can affect humans.
This research is expected to be a crucial stepping stone for future work in nutrition, genetics and medicine. Finding where and when the gene gets turned off is critical in determining which species are most likely to have the toxic sugar Neu5Gc, and if so what their use can be in transplants, diets, and research. The work of Alvarez-Ponce's team will make it easier to understand why certain diseases form and find ways to combat the spread of disease.
The results of the research will be published in the January issue of Oxford University Press science journal Genome Biology and Evolution.
More information: Sateesh Peri, Asmita Kulkarni, Felix Feyertag, Patricia M. Berninsone, David Alvarez-Ponce. "Phylogenetic distribution of CMP-Neu5Ac hydroxylase (CMAH), the enzyme synthetizing the pro-inflammatory human xeno-antigen Neu5Gc". Genome Biology and Evolution, doi.org/10.1093/gbe/evx251
Provided by University of Nevada, Reno