Researchers create 3-D cell arrays for more realistic experimental biological environments
Paving the way for testing experimental drugs in more realistic environments, scientists at the National Institute of Standards and Technology (NIST) have discovered how to make tiny colonies of cells grow in useful new ways inside petri dishes.
The research team's discoveries might help designers of miniature "lab-on-a-chip" technologies to grow three-dimensional colonies of liver cancer cells inside a chip's tiny chambers, rather than the merely two-dimensional colonies that they generally can culture now. Since many solid-tissue tumors are themselves three-dimensional, 3-D cell arrays could furnish more realistic biological environments for testing pharmaceuticals than those that are currently available.
"As tumor cell lines are commonly used for testing anti-cancer compounds, the biomedical community is actively looking for ways to test these drugs in 3-D cell cultures," said Darwin Reyes-Hernandez, a biomedical engineer at NIST. "Our findings could help bridge the gap between analyses of cells in the lab and in living creatures, a gap that currently limits the drug discovery processes."
To fulfill the promise of lab-on-a-chip technology, the chip's interior needs to have many features in common with the body itself, such as many different cell types growing in each other's presence. Scientists can already explore what a single cell type would do in the presence of a drug molecule simply by growing them together in a laboratory petri dish. But drugs must work in the body, not just a lab experiment. To study cell-cell interactions in a controlled fashion, scientists grow multiple cell types in the dish, making each type grow in a different location by changing the characteristics of the growing surface, a technique called micropatterning.
The NIST team, whose members specialize in the microfluidic technologies that would form much of the lab on a chip's physical environment, initially had the goal of making two different types of human cells grow side by side on a surface: liver cancer cells as well as endothelial cells, which line blood vessels in the body and are critical for cancer progression. Merely finding a way to create this shared boundary between two cell types would have been a worthy accomplishment, according to team member Kiran Bhadriraju, a NIST guest researcher who is visiting from Theiss Research in La Jolla, California. Existing technologies to create such micropatterned co-cultures are cumbersome, he said, and not easily used in large-scale pharmaceutical testing.
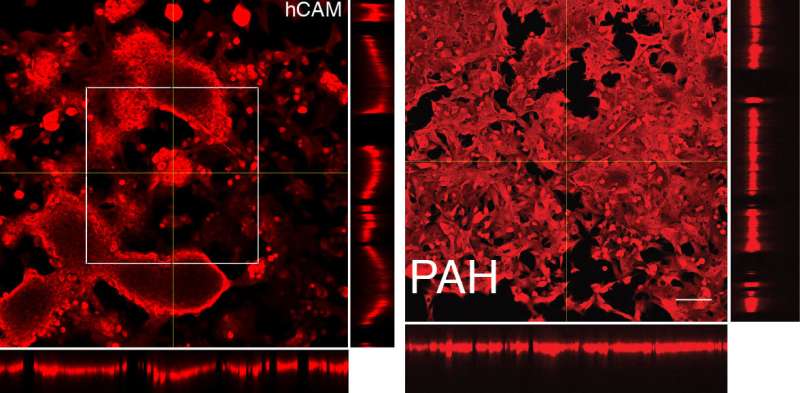
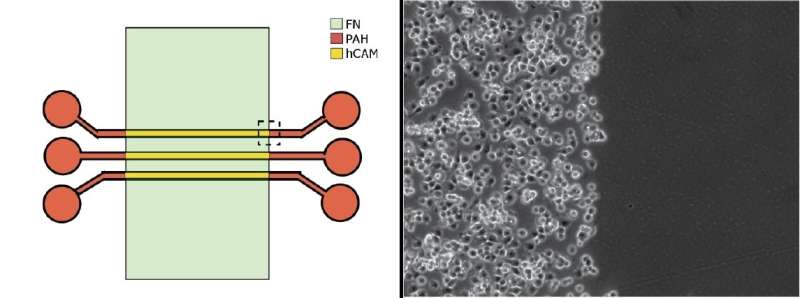
The team theorized that when they coated the surface with two different adhesives—fibronectin alone and a composite of fibronectin and other substances called hybrid cell adhesive material (hCAM)—the liver cancer cells would readily stick only to the hCAM, while the endothelial cells would adhere to the fibronectin. Preliminary experiments validated their hunch, and the discovery provided the NIST scientists with a way to create co-cultures of the tumor and endothelial cells where they wanted them.
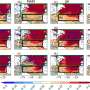
Creating the shared boundary they'd initially sought was an accomplishment on its own, but there was more to come. When they took images of the cells using a technique known as laser confocal microscopy, the team also discovered that the cells on the hCAM surface had grown layered arrays in three dimensions. Adding a third protein called transglutaminase—a sticky enzyme that glues protein molecules together—they could make the liver cancer cells instead form arrays only a single cell thick, giving them control over the process.
Knowing this relatively simple relationship among the chemicals, surface and liver cancer cells could be useful for culturing cancer cells together with completely different cell types, he says, and might allow these small cell cultures to be scaled up for the sort of high-throughput work a drug company would need to test large numbers of drug candidates.
"We expect that other cancer cell lines can be used for micropatterning similar co-cultures," said Bhadriraju. "While the liver cancer cell line used here is an important cell line for the pharmaceutical industry for testing anti-cancer drugs, we haven't tested yet whether other cancer cell types will form the same types of 3-D structures. But we're optimistic, as these proteins we coated the surface with are commonly used with other kinds of cancer cells."
More information: Kiran Bhadriraju et al. Fibronectin in Layer-by-Layer Assembled Films Switches Tumor Cells between 2D and 3D Morphology, ACS Biomaterials Science & Engineering (2017). DOI: 10.1021/acsbiomaterials.7b00608
Provided by National Institute of Standards and Technology
This story is republished courtesy of NIST. Read the original story here.