Non-steady state mass action dynamics reconsidered
The law of mass action was first described 150 years ago. Today, it is still a staple concept in every first-year chemistry text. The law states that the rate of a chemical reaction is proportional to the concentration of the reactants. For solutions in dynamic equilibrium, the law predicts how those solutions will behave.
But it is a challenge to apply the law of mass action to complex systems, such as biological metabolism. For one, most rate constants are not available, and measuring the missing ones is very labor-intensive.
In biological research, measuring rate constants requires isolating and purifying individual enzymes, then determining the rate constants with spectrophotometry - that is, by how much light a chemical absorbs. This slows down throughput speeds, making it hard to characterize each new genome. After all, to do this requires thousands of rate parameters.
Because of these issues, a large number of enzyme rate parameters have been determined for just a few well described systems, such as Escherichia coli. For most biological systems, scientists have not yet obtained rate constants.
Meanwhile, the method of choice for modeling large-scale biological processes (such as metabolism) involves constraint-based flux models. But these models suffer from a lack of physics regarding the law of mass action, limiting the ability of constraint-based approaches even when they are extended to include thermodynamics and other factors.
In answer, some researchers have proposed studying elementary reactions using the law of mass reaction through reaction affinities instead of rate constants.
A new paper in the journal Physical Biology by William R. Cannon and Scott E. Baker at the Pacific Northwest National Laboratory reviews the relationship between reaction affinity and free energy for isolated reactions by using concepts from statistical thermodynamic integration. Cannon and Baker set out to explicitly demonstrate the relationship between kinetic formulations of rates that use rate constants and a statistical thermodynamics formulation that uses chemical potentials.
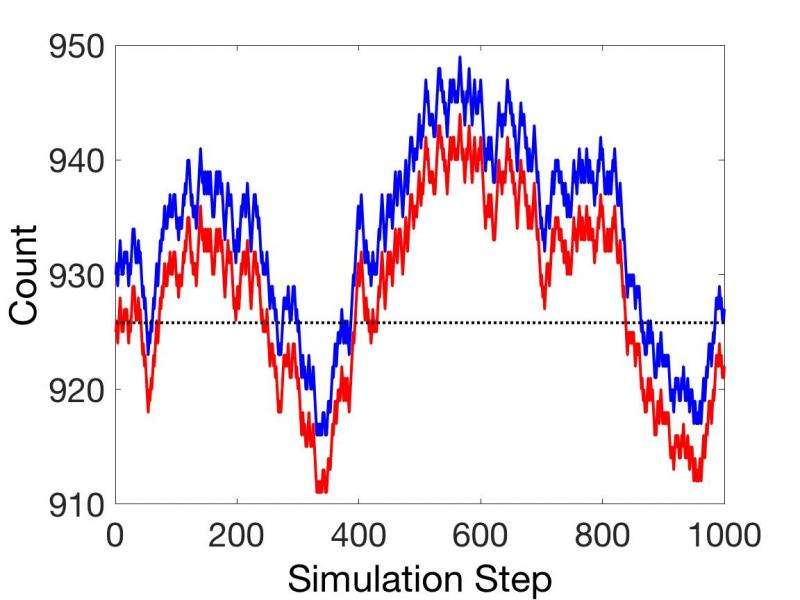
Subsequently they present a theorem for coupled reactions based on chemical potentials, which can provide relative rates for all sequentially coupled reactions under all conditions, steady state or not. This obviates the need for rate constants in many applications.
Using chemical potentials instead of rate constants represents a methodological leap that the authors say could signal a major advance for predictive modeling in systems biology.
The new method will enable complex, large-scale biological systems such as metabolism to be modeled with kinetics and thermodynamics. This will result in more accurate predictions for the design of microorganisms to behave in desired ways, such as those that could be harnessed to produce biofuels.
Moreover, the authors say, the new method will shed fundamental insight into the thermodynamic coupling between living organisms and their environment.
More information: William R Cannon et al. Non-steady state mass action dynamics without rate constants: dynamics of coupled reactions using chemical potentials, Physical Biology (2017). DOI: 10.1088/1478-3975/aa7d80
Journal information: Physical Biology
Provided by Pacific Northwest National Laboratory