An evolutionary breakpoint in cell division
Japanese researchers from Osaka University have discovered that the interaction between two proteins, M18BP1/KNL2 and CENP-A, is essential for cell division in various species except for mammals including human.
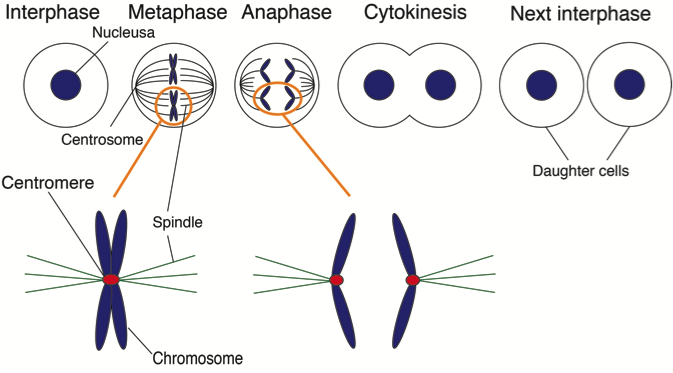
Mitosis is a process in living organisms in which a cell divides into two new daughter cells. It is necessary for growth, and replacement and repair of older cells. During mitosis, all genome information in an individual cell is copied and then divided equally into the daughter cells.
The centromere is a critical structure consisting of DNA and proteins that plays an important role in the distribution of genome information during mitosis. If the centromere is disrupted, an unequal division of the genome information can occur, resulting in defective cells and possibly cancer. Specifically, Osaka University Professor Tatsuo Fukagawa, who was the principal investigator of the study, was interested in mechanisms how the centromere is established and maintained in the cells.
"CENP-A is a key factor for the establishment and maintenance of the centromere," he said. "We looked into the mechanisms how CENP-A is deposited into centromere and especially focused relationship between the Mis18 complex and CENP-A, because previous studies had shown it regulates CENP-A deposition to the centromere."
"Research has suggested that the Mis18 complex associates with CENP-A on chromatin (old CENP-A) for new CENP-A deposition in non-mammalian vertebrates like chicken, but this unique loading system is lost in mammalian cells" he added.
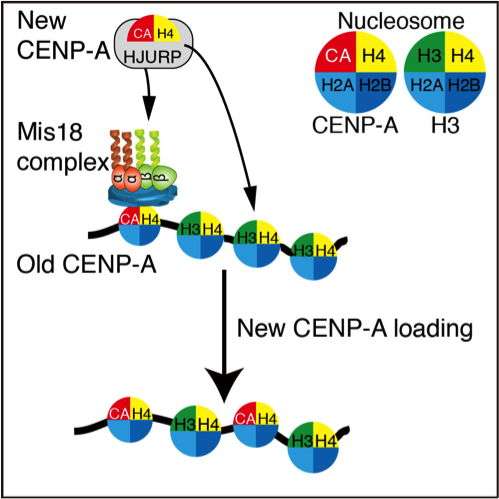
The Mis18 complex is made up of three subunits: Mis18a, Mis18b, and M18BP1/KNL2. Fukagawa and his team found that the M18BP1/KNL2 subunit in chicken had a specific binding motif that allowed the complex to bind to CENP-A on chromatin, which allowed new CENP-A deposition into the centromere to maintain centromeres in chicken cells.
Interestingly, evidence suggests a similar mechanism in plant and frog cells, but not in human or mouse cells.
"We predict M18BP1/KNL2 does not bind CENP-A in human or mouse cells. This binding motif is lost during evolution," said Fukagawa.
"When we looked further into the relationship of M18BP1/KNL2 with CENP-A, we found that CENP-A nucleosomes bind to M18BP1 through a CENP-C-like motif. It is really interesting to know why mammalian cells lose such an efficient mechanism"
As a next question, Fukagawa would like to determine the mechanisms how other centromere proteins on CENP-A nucleosomes are assembled to form functional centromeres, which are common structure in all organisms.
More information: Tetsuya Hori et al. Association of M18BP1/KNL2 with CENP-A Nucleosome Is Essential for Centromere Formation in Non-mammalian Vertebrates, Developmental Cell (2017). DOI: 10.1016/j.devcel.2017.06.019
Journal information: Developmental Cell
Provided by Osaka University