Researchers identify new mechanism for keeping DNA protein in line
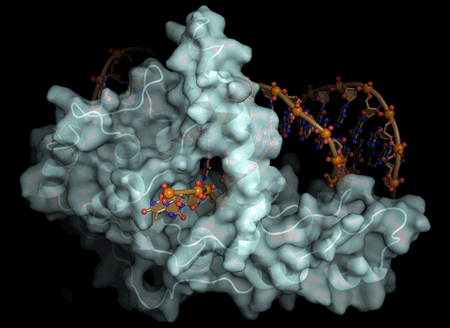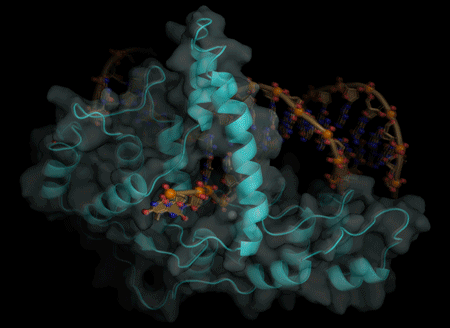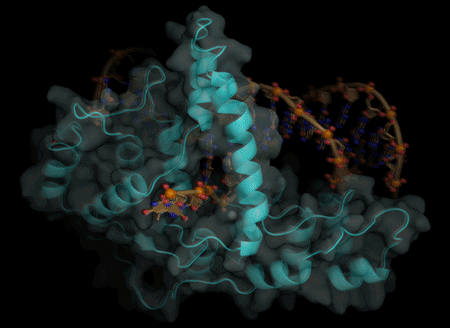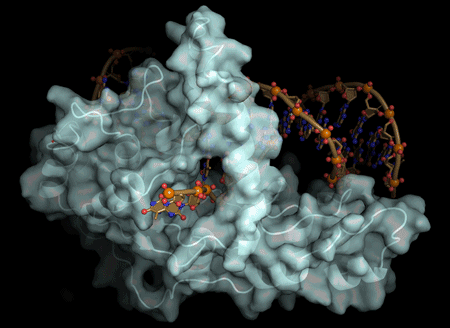
The actions of a protein used for DNA replication and repair are guided by electrostatic forces known as phosphate steering, a finding that not only reveals key details about a vital process in healthy cells, but provides new directions for cancer treatment research.
The findings, published this week in the journal Nature Communications, focus on an enzyme called flap endonuclease 1, or FEN1. Using a combination of crystallographic, biochemical, and genetic analyses, researchers at the Department of Energy's Lawrence Berkeley National Laboratory (Berkeley Lab) showed that phosphate steering kept FEN1 in line and working properly.
"FEN1, like many DNA replication and repair proteins, have paradoxical roles relevant to cancer," said study lead author Susan Tsutakawa, a biochemist at Berkeley Lab's Molecular Biophysics and Integrated Bioimaging Division. "A mistake by FEN1 could damage the DNA, leading to the development of cancer. On the other side, many cancers need replication and repair proteins to survive and to repair DNA damaged from cancer treatments. New evidence shows that phosphate steering helps ensure that FEN1 behaves as it should to prevent genome instability."
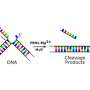
During the process of replication, double-stranded DNA unzips to expose the nucleotides along its two separate strands. In that process, flaps of single-stranded DNA are created. The job of FEN1 is to remove those flaps by positioning metal catalysts so that they can break down the phosphodiester bonds that make up the backbone of nucleic acid strands. This cleavage action occurs in the duplex DNA near the junction with the single-stranded flap.
Flaps that remain uncleaved can lead to toxic DNA damage that either kill the cell or cause extensive mutations. For example, trinucleotide repeat expansions, a mutation associated with disorders such as Huntington's disease and fragile X syndrome, are characterized by the failure of FEN1 to cut off the excess strand.
"What had been unclear before our study was how FEN1 was able to identify its exact target while preventing the indiscriminate cutting of single-stranded flaps," said Tsutakawa. "There must be a way for this protein to not shred similar targets, such as single-stranded RNA or DNA. Getting that right is critical."
Tsutakawa worked with corresponding author John Tainer, Berkeley Lab research scientist and a professor at the University of Texas, at the Advanced Light Source, a DOE Office of Science User Facility that produces extremely bright X-ray beams suitable for solving the atomic structure of protein and DNA complexes. Using X-ray crystallography, they were able to get a molecular-level view of the FEN1 protein structure.
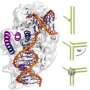
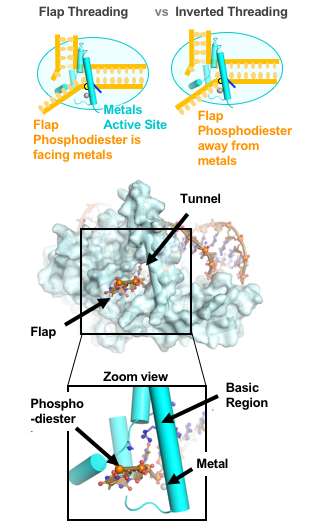
They determined that the single-stranded flap threaded through a small hole formed by the FEN1 protein. The size of the hole serves as an extra check that FEN1 is binding the correct target. However, they surprisingly found that the single-stranded flap is inverted such that the more vulnerable part of the DNA, the phosphodiester backbone, faces away from the metal catalysts, thereby reducing the chance of inadvertent incision.
The inversion is guided by a positively charged region in FEN1 that stabilizes the upside-down position and steers the negatively charged phosphodiester of the single-stranded DNA through the FEN1 tunnel.
"These metals are like scissors and will cut any DNA near them," said Tsutakawa. "The positively charged region in FEN1 acts like a magnet, pulling the flap away from these metals and protecting the flap from being cut. This is how FEN1 avoids cutting single-stranded DNA or RNA."
"This phosphate steering is a previously unknown mechanism for controlling the specificity of FEN1," she added. "Cancer cells need FEN proteins to replicate, so understanding how FEN1 works could help provide targets for research into treatments down the line."
More information: Susan E. Tsutakawa et al, Phosphate steering by Flap Endonuclease 1 promotes 5′-flap specificity and incision to prevent genome instability, Nature Communications (2017). DOI: 10.1038/ncomms15855
Journal information: Nature Communications
Provided by Lawrence Berkeley National Laboratory