Researchers send DNA on sequential building mission
A team of scientists has developed a method to create structures whose building blocks are a millionth of a meter in size by encoding DNA with assembly instructions.
The work, described in the journal Nature Communications, manipulates the sequencing of DNA to offer an intricate and innovative approach to synthesize materials at the most fundamental level.
"Sequential programmability is a powerful addition to the self-assembly toolbox that will prove useful in creating the tiniest of materials," explains Yin Zhang, the paper's lead author and a graduate student at New York University's Center for Soft Matter Research. "It brings some of the advantages of the biological use of controlled sequential assembly using molecules, nucleic acids, and proteins to a new design scale."
NYU physics professors Paul Chaikin and Jasna Brujic as well as Nadrian Seeman, an NYU professor of chemistry, co-directed the research.
Both natural and human-made structures are built sequentially—from cells to skyscrapers. Like Russian nesting dolls, assembly takes place on the inside before commencing on the outside.
However, when making materials on a micrometer scale, or about one hundredth of the width of a strand of human hair, scientists face challenges unfamiliar to engineers and manufacturers.
While many methods have been adopted to manipulate such tiny particles, these approaches all have notable shortcomings in assembling structures.
The NYU team sought to overcome these hurdles with a new approach: encode the instructions of assembly within the building blocks and let these building blocks self-organize into a prescribed structure in a pre-determined sequence.
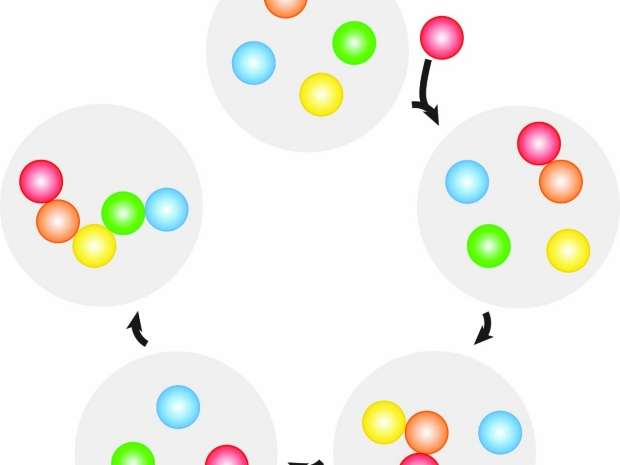
To do so, it deployed different strands of DNA, each coated on a droplet of oil in water, where they then "talked" to each other through DNA-mediated interactions. Specifically, the scientists placed four "flavors" of droplets—labeled B, C, D, and E—into water. They then added an "initiator" droplet, A, which began the sequencing process. Here, the DNA strand on A initiates a chain of events in which it displaces one of the paired strands on B, whose released strand moves to activate C, the next droplet in the sequence, and so on. The process results in a droplet chain, ABCDE.
Journal information: Nature Communications
Provided by New York University