Biosynthetic secrets: How fungi make bioactive compounds
Biological engineers at Utah State University have successfully decoded and reprogrammed the biosynthetic machinery that produces a variety of natural compounds found in fungi.
Fungi and bacteria produce a range of bioactive natural products that exhibit anti-cancer, anti-microbial, herbicidal, insecticidal and anti-cholesterol properties, among others.
These useful molecules, known as nonribosomal peptides, are formed by a group of enzymes called nonribosomal peptide synthetases (NRPSs) that assemble a diverse group of natural products including penicillin (antibacterial), beauvericin (anticancer) and vancomycin (antibacterial). While bacterial NRPSs are well understood, the fungal versions of these processes are only now coming to light.
Dr. Jixun Zhan, a professor of biological engineering at USU, studies the catalytic synthesis of natural products in bacteria and fungi. He and his team have reproduced many bio-active compounds in engineered microbes. Most recently, the team biosynthesized beauvericin and bassianolide—natural compounds originally produced by the fungus Beauveria bassiana that are known to have multiple beneficial effects.
The team's findings, published May 23 in Nature Communications, are the first to describe the difference between bacterial and fungal iterative NRPS mechanisms. Zhan says a clearer understanding of NPRS processes could lead to advances in synthesizing existing and new compounds for drug discovery.
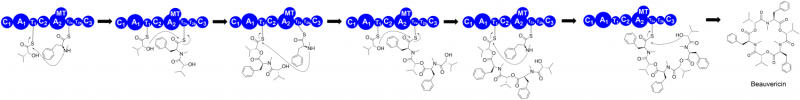
"Our goal in this research was to understand how fungal iterative NRPSs precisely and repetitively use their catalytic units to make these different compounds," said Zhan. "We're specifically looking at the synthetases that assemble beauvericin and bassianolide and trying to understand how they are assembled and how the NRPSs are programmed to control molecule length and other factors."
NRPSs are modular enzymes that contain a series of small catalytic units called domains. Zhan's team studied two fungal NRPSs from B. bassiana that are responsible for the synthesis of the two compounds.
The researchers successfully cut the large enzymes into functional fragments and reconstituted their activity in baker's yeast. Through a combination of enzyme dissection, domain swapping, site-directed mutagenesis and in vitro enzymatic reactions, the team has revealed fungal NRPSs' chain elongation and length control strategy.
More information: Dayu Yu et al, Decoding and reprogramming fungal iterative nonribosomal peptide synthetases, Nature Communications (2017). DOI: 10.1038/NCOMMS15349
Journal information: Nature Communications
Provided by Utah State University