Using nature to build nanomachines
Flagella are tiny rotary motors that move bacteria. A research team led by Keiichi Namba, professor at Osaka University has used electron cryomicroscopy to show that a small difference of only several amino acids can have a large impact on flagella function. The study gives new insight on the construction of synthetic nanomachines and can be read in Nature Communications.
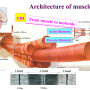
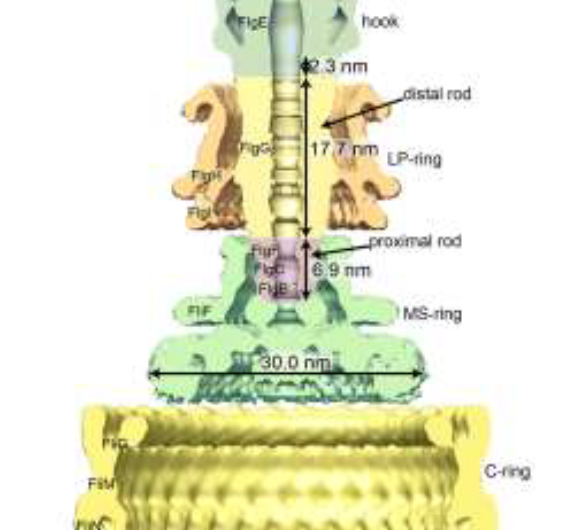
The flagellar motor can be subdivided into three subunits that include the helical propeller (filament), universal joint (hook), and drive shaft (rod). The hook is flexible and transmits torque to the filament, while the rod is remarkably stiff and can rotate >1000 Hz in some bacteria. The hook is 55 nm long and 18 nm wide, and the rod is 20 nm long and 13 nm wide. Structurally, the hook consists of one protein, FlgE, while the rod consists of several proteins, but only FlgG is in direct contact with the hook. FlgE and FlgG include two domains that are thought responsible for the very different mechanical properties of the rod and hook even though in many ways the domains are almost identical.
"The different functions can be explained by how the domains are packed," said Namba, whose lab studies nanomachines in nature, adding that, "In the rod, they are highly packed, which makes the rod rigid. In the hook, the domains are spaced out, giving the hook flexibility."
High-resolution electron cryomicroscopy images and corresponding atomic models suggested the different packing is related to a slightly different tilt of 7o in the orientation of the domains. This 7o tilt reduces the packing, which allows the hook protofilament to compress and expand, explaining its flexibility.
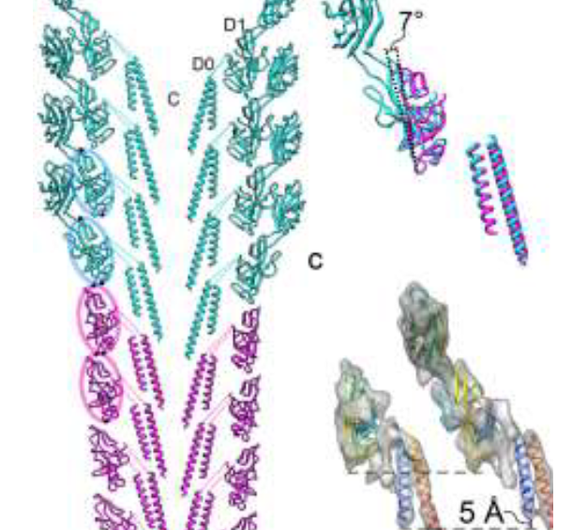
Namba noted that although similar, one particular part the two proteins is longer in FlgG by 18 amino acids. To confirm that these amino acids determined the rigidity of the rod, his team inserted them to the hook protein and found that the hook became much stiffer.
"The 18 amino acid insert shows how a small changes can drastically change the behavior of a nanomachine," he said.
Understanding the structural design of the flagellar motor is expected to provide a blueprint for the synthetic construction of similar nanomachines.
More information: Takashi Fujii et al, Identical folds used for distinct mechanical functions of the bacterial flagellar rod and hook, Nature Communications (2017). DOI: 10.1038/ncomms14276
Journal information: Nature Communications
Provided by Osaka University