Cultivated scallops populations develop distinct genetic structure
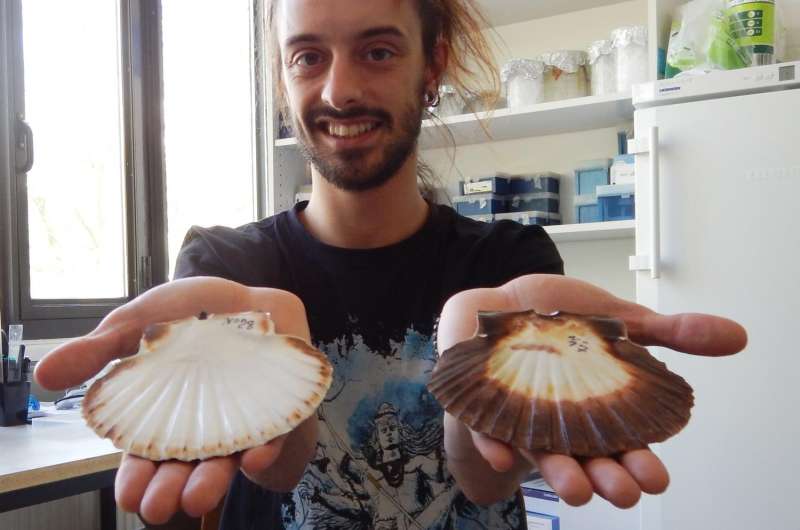

The scallop is one of the largest edible molluscs, and gourmets consider it to be a great delicacy. To meet this demand, the fishing industry cultivates these shellfish in coastal aquafarms. In a new analysis, behavioural ecologists at Bielefeld University have confirmed that cultivated scallops developed their own genetic structure that differs from that of natural scallops. The biologists studied a total of nine populations of scallops (Pecten maximus) along the coast of Northern Ireland. They are presenting their results this Wednesday in the research journal 'Royal Society Open Science'.
'Of the nine scallop populations studied, only one shows a marked genetic difference from the others and that is the artificially cultivated type,' reports Joseph I. Hoffman, head of the Molecular Behavioural Ecology research group. New breeds are cultivated in, for example, mesh cages in coastal waters. Now and then, young scallops escape through the mesh and are thereby able to impact on natural populations. Biologists use the term population to describe a group of organisms of one species that live together in one area and are linked together genetically through reproduction over successive generations.
The researchers analysed the genetic architecture of the mollusc populations. 'Studying the genetic architecture of animal populations helps us to understand which external appearance an organism can adopt - for example, how large a mollusc can become or whether it can develop red streaks on its surface,' says David Vendrami. The doctoral student has analysed a total of 180 mollusc samples. The Agri-Food and Biosciences Institute in Belfast (Northern Ireland) collected these in February 2015 during an excursion along the Northern Irish Atlantic coast.
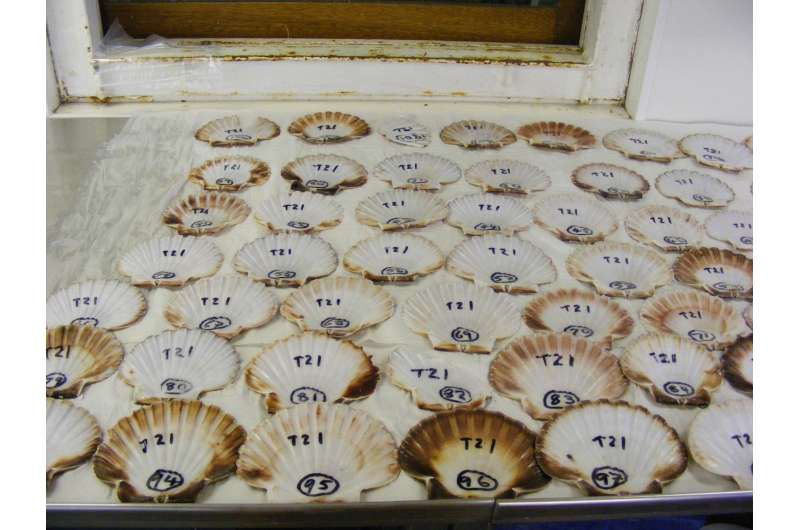

The researchers have not just confirmed how breeding affects scallop populations. Their study also confirms that these molluscs adapt their shape and internal colouring very flexibly to conditions in their environment, and that they do this independently of whether they belong to the one cultivated or the eight natural populations. 'We have tested how far genes relate to appearance. However, that is very probably not the case. It is highly likely that the external characteristics of the molluscs depend on their surroundings,' says Vendrami.
Scientists at Bielefeld also used this study to compare a classical DNA analysis with a new procedure. The classical analysis proceeds by evaluating repeated short DNA sequences (microsatellites) in order to compare samples from different organisms. The modern procedure (RAD sequencing) takes less time to analyse thousands of times more DNA sequences. 'The new procedure is markedly better at finding differences in the populations than the classical approach,' says David Vendrami.
In their future research, Hoffman, Vendrami, and their colleagues will be going beyond Northern Ireland and studying samples along the entire Atlantic coast from Norway to Portugal as well as in the Mediterranean. Their aim is to find out how scallops as well as other crustaceans react to different environmental conditions in terms of their growth.
More information: David L. J. Vendrami et al, RAD sequencing resolves fine-scale population structure in a benthic invertebrate: implications for understanding phenotypic plasticity, Royal Society Open Science (2017). DOI: 10.1098/rsos.160548
Journal information: Royal Society Open Science
Provided by Bielefeld University