Tuberculosis virulence factor identified, may be target for new drug
Scientists have discovered the mechanism that hijacks the immune system's response to tuberculosis, revealing an important new drug target for the disease that kills more than 1 million people each year.
Herman Sintim, Purdue University's Drug Discovery Professor of Chemistry, collaborated with scientists at Johns Hopkins University to determine how tuberculosis turns off a human cell's signal to mount an immune response to the bacteria. Their findings were published in the journal Nature Chemical Biology.
Tuberculosis is a bacterial disease that results in coughing, fever, night sweats, weight loss and sometimes death.
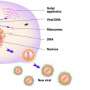
When Mycobacterium tuberculosis enters a human cell, the presence of its DNA and a molecule that it makes called c-di-AMP alert the cell to the bacteria's presence. The human cell responds by creating a messenger molecule, cGAMP, which signals nearby cells to mount an immune response to kill the tuberculosis bacteria.
The human cell also produces another molecule, ENPP1, which degrades the cGAMP. That key step turns off the call for an immune response.
"Immune response can involve reactive oxygen and nitrogen species, which can kill the bacteria but at the same time cause collateral damage and also damage or kill the host cells as well," Sintim said. "There is a very delicate response to bacteria and stopping that response once bacteria have been taken care of."
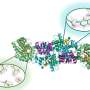
But the tuberculosis bacterium has found a way to turn off the call for help. By producing a protein called cyclic dinucleotide phosphodiesterase (CdnP), the bacterium reduces the concentration of the cell's messenger molecule, cGAMP, a nucleic acid. This accentuates the effect of the human phosphodiesterase ENPP1, an enzyme that cleaves nucleic acids, to quickly degrade any already-made cGAMP and turn off the immune response early.
"The host cGAMP never gets to a high enough concentration to activate the immune response," Sintim said. "This is a very effective strategy the bacteria have developed to suppress an immune response."
Sintim and colleagues tested their hypothesis by creating a mutant of Mycobacterium tuberculosis that lacked the CdnP protein and tested it in a mouse model. On average, the mice with the mutant bacteria lived more than two times longer than mice with the wild type, suggesting that CdnP played a role in suppressing immune response.
They then artificially synthesized the cGAMP molecule and investigated if it was a substrate for CdnP. The CdnP degraded the human molecule as predicted.
Sintim said the CdnP protein in the tuberculosis bacteria now becomes an attractive target for a new drug. If a molecule could be developed that would inactivate or inhibit CdnP, it would improve immune response in tuberculosis patients.
Sintim's team identified several molecules that would bind with and inhibit CdnP, but they have not reached the potency level needed to create drugs. They will continue looking for new compounds that could potently inhibit this newly discovered CdnP drug target.
More information: Ruchi Jain Dey et al. Inhibition of innate immune cytosolic surveillance by an M. tuberculosis phosphodiesterase, Nature Chemical Biology (2016). DOI: 10.1038/nchembio.2254
Journal information: Nature Chemical Biology
Provided by Purdue University