An ancient mechanism helps a cell to resist stress
Biologists from the Lomonosov Moscow State University in collaboration with colleagues from the Engelhardt Institute of Molecular Biology, Russian Academy of Sciences, used RNA transfection and in vitro techniques to show how the same mRNA can direct protein synthesis in a cell by four different means. The research results have been published in a peer-reviewed journal Scientific Reports.
Scientists from the Belozersky Institute of Physico-Chemical Biology, a department of the Lomonosov Moscow State University, along with their colleagues, have applied a transfection method to deliver RNA into the cell to observe the impact of cell stress on protein biosynthesis on a short-time scale.
Cell stress and reprogramming of protein synthesis
Sergey Dmitriev, senior researcher at the Belozersky Institute of Physico-Chemical Biology, the Lomonosov Moscow State University, lead author of the article, says, "Our project is devoted to the studies of protein biosynthesis mechanisms, including the situations of cell stress. The research highlights three aspects. The first one concerns methods as we present a technique, which allows analyzing protein synthesis in a cell with the help of a short-term RNA transfection technique".
Transfection is a method of DNA and RNA delivery to a living cell. Usually, DNA is used. After introduction into the nucleus, it initiates the processes of new RNA synthesis, and only afterwards are the RNAs exported into the cytoplasm to participate in protein production. Biologists from the Lomonosov Moscow State University propose a methodology of introducing an artificially synthesized RNA into the cell as a template for immediate protein synthesis.
RNA is delivered to cells with the help of a special chemical agent. Once introduced into the cytoplasm, it participates in protein production on contact with a ribosome, a much shorter process. In as little as one to two hours, the researchers observed protein activity and estimated its quantity.
This technique allows studying the impact of stress on the cell over a short time scale. Cell stresses include, for example, heat shock caused by elevated temperature; oxidative stress provoked by reactive oxygen species; and response to chemical agents that disrupt homeostasis (including antibiotics and medical drugs). Factors of cell stress compel the cell to suspend protein biosynthesis (or "reprogram" it), until the system redresses the balance.
Sergey clarifies: "Usually, these processes last from one to four hours, and our technique of "fast" RNA transfection is the most convenient way to study the effect of these processes. We've conducted our research on cultured human kidney cells, which serve as a standard model for such studies. Finally, we have elaborated a technique that allows obtaining artificial RNA, transfered it into cells and obtained the result in a very short period of time. We've named the whole method FLERT (for fleeting mRNA transfection), which sounds in Russian a bit like 'flirtation,'" says Sergey.
Why is the sum of 40S and 60S equal to 80S in case of a ribosome?
Messenger RNA (mRNA) is a polymer of nucleotides coding for a protein. One amino acid is encoded by three nucleotides. There is a special molecular machine, the ribosome, that produced protein. Moving along mRNA, the ribosome reads information in a triplet-by-triplet manner.
The structure of the protein synthesis machine is very complicated. It comprises two subunits— a small one (40S) and a large one (60S). A whole ribosome is obtained when they join. However, it's specified not as 100S, but as 80S. The reason is that these figures refer not to the particle mass, but to the sedimentation coefficient, determined during centrifugation. This coefficient depends on several parameters, including the shape of a particle.
In order to start information decoding, first of all it is necessary to find the right starting point—a triplet from which the reading begins. Detection of the starting point is problematic, as there are no marks for triplets in mRNA. However, if you start reading from the wrong nucleotide, the reading frame will be shifted, and everything will go wrong. Special proteins (translation initiation factors) help the ribosome to find the right place in the template to start reading triplets.
Usually, there is a distance between the beginning of the mRNA chain and the starting point, called "leader." A ribosome should pass this leader by without reading. The scientists wondered what would happen if mRNA began right with the starting codon—i.e., from the "first word." It's interesting that in archaea (single-celled prokaryotic organisms that have been living on the Earth for billion years and are capable of surviving in extreme conditions) and some other primitive organisms, most mRNAs begin right from the start codon. Such RNAs are called "leaderless." Leaderless mRNAs are supposed to be an evolutionary prototype of messenger RNAs because ancient ribosomes were unable to find starting points and initiated decoding from the very beginning of the mRNA chain.
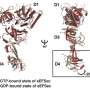
A ribosome passes through several phases in order to interact with mRNA and start protein synthesis. Normally, the 40S subunit of the ribosome binds mRNA, and then the large 60S subunit joins it at the start codon. In contrast, the leaderless mRNA can be loaded directly into the whole ribosome. This discovery was made in the 1990s by Ivan Shatsky, professor at the Lomonosov Moscow State University.
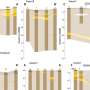
In the new project, scientists have demonstrated that due to their unique properties, leaderless RNAs are resistant to many stress types and continue directing protein synthesis even in such conditions, while common RNAs with leaders stop working in the first minutes after the impact. Using the FLERT technique, scientists have now demonstrated this in living cells.
The research extension has brought even more interesting results. It turns out that unique properties of the leaderless mRNA provide it with high flexibility in the choice of protein synthesis mechanisms. The researchers discovered that eukaryotes possess several pathways by which a ribosome could find itself on the start codon. These modes are mediated by distinct sets of specialized proteins called translation initiation factors, and have been shown to operate on different mRNAs.
The most common pathway, which could be used by any cellular mRNA, is provided by the eIF2 protein. However, this factor is very quickly inactivated under conditions of stress. As a result, ribosomes fail to recognize the start codons on all mRNAs except those that use other initiation factors.
In a previous study, the scientists discovered that eIF2 is not the only factor able to do this work. For instance, mRNA of the hepatitis C virus is capable of doing without eIF2 and can use eIF5B or eIF2D instead. This virus was supposed to be unique in this sense—while canonical templates are passively waiting until a ribosome binds them, the hepatitis C virus mRNA "grasps" the 40S subunit and puts it in the right place on the chain. And now scientists have proved that the leaderless mRNA is capable of acting in the same way.
It's also interesting that all organisms possess eIF5B factor, as it's a conserved evolutionary protein. In contrast, eIF2 exists only in eukaryotes and archaea—so it's not universal. All the above-mentioned results suggest that the well-studied classical factor eIF2 is needed only if ribosomes recognize mRNA by active searching for the start codon. Such means of translation initiation is called "scanning," and requires eIF2. When the start codon is found, eIF2 is replaced by eIF5B and protein synthesis starts. Ancient leaderless mRNA can use a primitive mechanism, immediately recruiting eIF5B factor.
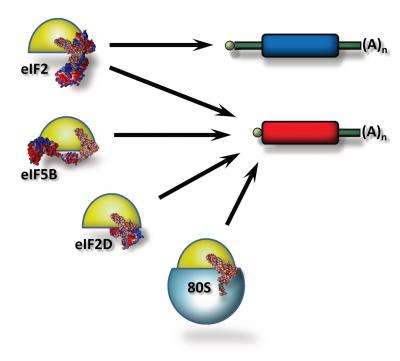
Sergey Dmitriev concludes: "We've got a nice result explaining everything. We've found out that a primitive mRNA could use an ancient evolutionary mechanism. Moreover, it is capable of using the other three pathways—through eIF2, eIF2D or direct recruitment of the whole 80S ribosome."
More information: Kseniya A. Akulich et al, Four translation initiation pathways employed by the leaderless mRNA in eukaryotes, Scientific Reports (2016). DOI: 10.1038/srep37905
Journal information: Scientific Reports
Provided by Lomonosov Moscow State University