Proximity labeling reveals the key components of a structure that gives cells their sense of place
The protein complexes cells use to attach to the local biological matrix do more than hold cells in place, they help the cell sense what tissue they're in and what cell type they should be. However, the unstable nature of these 'focal adhesion' protein complexes makes them difficult to study. Researchers at A*STAR have built a working model of the focal adhesion by using a molecular tagging technique to precisely identify all the proteins involved1.
With an average cell containing 10-15,000 proteins, it is difficult to ascertain how specific proteins come together to perform certain functions. Paxillin is a well-known focal adhesion protein, but a "bewildering array" of other proteins have been proposed to make up the adhesion complex, says Ed Manser from the A*STAR Institute of Molecular and Cell Biology, who led the work.
"The problem is that proteomic methods are largely a smash and grab activity," Manser explains. "You can 'tag' a protein such as paxillin and grab it once the cell has been broken, in the hope of finding its cellular partners." His team takes a more careful approach: "we tag all the local proteins before we break the cell."
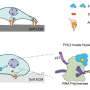
The 'BioID' tagging technique used by the team2, also developed at A*STAR, involves producing a cell line in which paxillin is fused to an enzyme that can add biotin to proteins within approximately 20 nanometers: an average protein is about five nanometers wide. These newly-biotinylated proteins are easily identified among the cellular rubble inside the ruptured cell.
The researchers combined BioID with stable isotope labeling to calculate the relative enrichment of proteins relative to paxillin and kindlin-2: they used this to infer their location within the focal adhesion structure.
Previous studies had proposed hundreds of proteins make up focal adhesions. The IMCB team identified just 35 proteins involved in in the process and suggest fewer than 50 distinct focal adhesion proteins. "We also confirmed that only seven of these proteins directly bind to paxillin, which is a credible number," Manser says.
The experiments also turned up a few surprises. Paxillin was previously thought to sit on the cell surface or 'plasma membrane'—but the lack of tagged local membrane proteins indicated it lies some distance away.
"The excitement is to develop a proteomic technique that can actually give you much better resolution than optical super-resolution methods," Manser says. Studies of other protein complexes are already underway.
More information: J.-M. Dong et al. Proximity biotinylation provides insight into the molecular composition of focal adhesions at the nanometer scale, Science Signaling (2016). DOI: 10.1126/scisignal.aaf3572
Kyle J. Roux et al. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells, The Journal of Cell Biology (2012). DOI: 10.1083/jcb.201112098
Journal information: Science Signaling , Journal of Cell Biology