Researchers identify new pathway in human pathogens
Several of the more aggressive pathogens that infect humans can thrive in an oxygen-free environment of the human gut. These pathogens also have the ability to acquire the essential nutrient iron from an abundant cofactor, specifically heme (the cofactor that makes blood and muscle appear red).
Newly published research from University of Georgia researchers reveals how a key enzyme in this new pathway functions to release the iron atom in the absence of oxygen. Further illumination of the atomic mechanism will provide an opportunity for a new class of antimicrobial compounds.
The study, "Radical new paradigm for heme degradation in Escherichia coli 0157:H7," was published in the early online edition of the Proceedings of the National Academy of Sciences on Oct. 10.
While the pathways for degrading of heme by pathogens to acquire iron in the presence of oxygen, or aerobically, have long been understood, the research presents new insights into how pathogens living in an anaerobic environment utilize heme as an iron source to survive. The discovery infers broad implications for research on pathogenic enzymes and especially those that achieve antibiotic resistance.
"There are decades of published research analyzing heme degradation, but every single pathway that has been investigated ultimately required molecular oxygen as a co-substrate in the degradation of heme," said William Lanzilotta, associate professor in the department of biochemistry and molecular biology in the Franklin College of Arts and Sciences and a corresponding author on the paper. "Shigella, cholera, and the hemorrhagic E. coli strains all have specific genes that allow them to outcompete the healthy microorganisms living in the anaerobic environment of the human gut. Our fundamental question was: How are they able to degrade the heme and liberate the iron without molecular oxygen?"
Bacteria evolve various enzymatic pathways for survival in the body, and the researchers narrowed their focus to three genes in the hemorrhagic E. coli that had not been characterized. The discovery also builds upon recent developments in what is known as radical SAM chemistry, a designation for a superfamily of enzymes that are very old, from an evolutionary perspective.
"These enzymes are primordial in that they utilize fundamental cofactors that have been on this planet for a very long time, going all the way back to anaerobic conditions on early Earth. Despite the simplicity of the cofactors, the chemistry is quite diverse and these anaerobic pathogenic organisms clearly take advantage of that," Lanzilotta said.
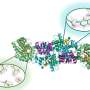
As the name implies, radical SAM enzymes utilize a radical intermediate in order to open the porphyrin ring in the absence of oxygen. The paper describes a possible mechanism for carbon-carbon bond breakage by a highly reactive radical intermediate.
"The radical SAM enzymes are present across all kingdoms of life, our body utilizes them to do some very sophisticated chemistry, in particular when it comes to things like bone formation," Lanzilotta said. "Because of the diversity of the chemistry, there are also many broader implications here. For example, other radical SAM enzymes are involved in the biosynthesis of new compounds with anti-biotic and anti-tumor properties as well as the production of commodity chemicals of great value."
Investigating the role of these enzymes in heme degradation could lead to the development of new strategies to fight pathogenic microorganisms, particularly among specific strains that have developed resistance to current anti-biotic treatments.
"Targeting this particular enzyme and this mechanism of anaerobic heme degradation would be a brand new strategy for developing new antimicrobial compounds that target aggressive pathogens," Lanzilotta said.
More information: Joseph W. LaMattina et al. Radical new paradigm for heme degradation inO157:H7, Proceedings of the National Academy of Sciences (2016). DOI: 10.1073/pnas.1603209113
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Georgia