Research on common bacterium opens door to fighting gastric cancer

A common bacterium that more than half of people have in their gut can use hydrogen gas present in the gastrointestinal tract to inject a cancer-causing toxin into otherwise healthy cells, according to a recently published study led by University of Georgia researchers.
The bacterium's reliance on hydrogen presents a pathway to potential new treatment and preventive measures in fighting gastric cancers, which kill more than 700,000 people per year, said corresponding author Robert Maier, Georgia Research Alliance Ramsey Eminent Scholar of Microbial Physiology in the Franklin College of Arts and Sciences.
Previous studies solidified the relationship between stomach ulcers and cancer and certain strains of Helicobacter pylori, a stomach-dwelling bacterium that causes 90 percent of all gastric cancers. Earlier research also found a link between a toxin known as CagA, or cytotoxin-associated gene A, and cancer formation, but the new study exposes how the bacterium uses hydrogen as an energy source to inject CagA into cells, resulting in gastric cancer, Maier said.
"There are many known microbes in the human gut that produce hydrogen and others that use hydrogen. The implications of the study are that if we can alter a person's microflora, the bacterial makeup of their gut, we can put bacteria in there that don't produce hydrogen or put in an extra dose of harmless bacteria that use hydrogen," Maier said. "If we can do that, there will be less hydrogen for H. pylori to use, which will essentially starve this bacteria out and result in less cancer."
Changing the microbial makeup of a person's gut sounds complicated, but scientists are already exploring ways to do so through the use of probiotics, antibiotics, nutritional regimes and even fecal transplants.
Although many people carry the H. pylori bacterium, it can take decades for the infection to develop into cancer, providing an opening for aggressive preventive measures in people who have a high risk of developing tumors, said lead author Ge Wang, a senior research scientist in the Department of Microbiology. The presence of H. pylori strains with both CagA and high hydrogen-utilizing activity within patients can potentially serve as biomarkers for predicting future cancer development.
An earlier study led by Maier, "Molecular Hydrogen as an Energy Source for Helicobacter pylori," published by the American Association for the Advancement of Science in 2002, showed that the presence of hydrogen in the gastric chamber was important for bacterial growth, but it wasn't until the current study that researchers learned of the strong cancer connection: The bacterium harnesses the energy from hydrogen gas, via an enzyme known as hydrogenase, to severely disrupt host cell function, leading to cancer.
"If hydrogen is in the gastric mucosa, of course a bacterium will use it," Maier said. "It's an excellent energy source for many bacteria out in nature. But we didn't realize that pathogens like H. pylori could have access to it inside an animal in a way that enables the bacterium to inject this toxin into a host cell and damage it."
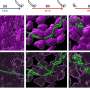
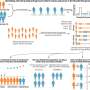
Not all strains of the bacteria cause cancer, but those with CagA are known carcinogens. Using human-derived gastric cells, Wang analyzed the hydrogenase activity in the cells that were infected with different strains—both cancer causing and non-carcinogenic—of H. pylori. He found more activity in the cancer-causing strains. After using genetic engineering to knock out the DNA fragment containing the hydrogenase-inducing genes, which prevented those strains from deriving energy from hydrogen, he found that the strain he created could no longer move the cancer-causing toxin into the gastric cells. Collaborators at Vanderbilt University took Wang's in vitro model and adapted it to test the theory in gerbils, which confirmed the UGA researchers' findings.
More information: Ge Wang et al. Hydrogen Metabolism inPlays a Role in Gastric Carcinogenesis through Facilitating CagA Translocation, mBio (2016). DOI: 10.1128/mBio.01022-16
Journal information: mBio
Provided by University of Georgia