Life as we know it most likely arose via 'long, slow dance'
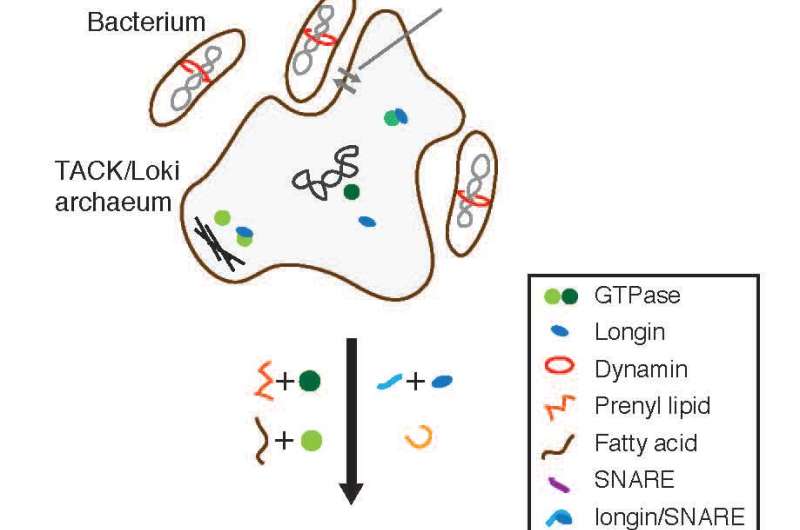
The first eukaryote is thought to have arisen when simpler archaea and bacteria joined forces. But in an Opinion paper published June 16 in Trends in Cell Biology, researchers propose that new genomic evidence derived from a deep-sea vent on the ocean floor suggests that the molecular machinery essential to eukaryotic life was probably borrowed, little by little over time, from those simpler ancestors.
"We are beginning to think of eukaryotic origins as a slow process of growing intimacy—the result of a long, slow dance between kingdoms, and not a quick tryst, which is the way it is portrayed in textbooks," says Mukund Thattai of the National Centre for Biological Sciences in India.
The eukaryotic cells of plants, animals, and protists are markedly different from those of their single-celled, prokaryotic relatives, the archaea and bacteria. Eukaryotic cells are much larger and have considerably more internal complexity, including many internal membrane-bound compartments.
Although scientists generally agree that eukaryotes can trace their ancestry to a merger between archaea and bacteria, there's been considerable disagreement about what the first eukaryote and its immediate ancestors must have looked like. As Thattai and his colleagues Buzz Baum and Gautam Dey of University College London explain in their paper, that uncertainty has stemmed in large part from the lack of known intermediates that bridge the gap in size and complexity between prokaryotic precursors and eukaryotes. As a result, they say, the origin of the first eukaryotic cell has remained "one of the most enduring mysteries in modern biology."
That began to change last year with the discovery of DNA sequences for an organism that no one has ever actually seen living near a deep-sea vent on the ocean floor. The genome of the archaeon known as Lokiarchaeum ('Loki' for short) contains more "eukaryotic signature proteins" (ESPs) than any other prokaryote. Importantly, among those ESPs are proteins (small Ras/Arf-type GTPases) critical for eukaryotes' ability to direct traffic amongst all those intercellular compartments.
The authors consider the available data to explore an essential question: what might the archaeal ancestor of all eukaryotes look like? "If we could turn back the clock and peer inside this cell, would its cellular organization have been like that of an archaeal cell or more eukaryote-like?" Dey says.
As the closest known archaeal relative of eukaryotes, Loki helps to answer that question. The researchers say that the ESPs found in Loki are unlikely to work in the same way they do in eukarytoes. That's because Loki doesn't appear to have enzymes required for ESP association with membranes or key building blocks of the membrane trafficking machinery.
"However," Baum says, "the genome can be seen as 'primed' for eukaryogenesis. With the acquisition of a number of key genes and lipids from a bacterial symbiont, it would be possible for Loki-type cells to evolve a primitive membrane trafficking machinery and compartmentalization."
The researchers predict that, when Loki is finally isolated or cultured, "it will look more like an archaeon than a proto-eukaryote and will not have internal compartments or a vesicle-trafficking network." But its morphology and/or cell cycle might have complexities more often associated with eukaryotes.
Baum and Dey say they now plan to explore the basic cell biology of the related archaea Sulfolobus acidocaldarius, first isolated from an acidic hot spring in Yellowstone National Park.
"We believe it will be very difficult to crack the mysteries of eukaryogenesis without first understanding the archaeal cell biology," Dey says. "We are currently developing tools in the lab to study the cell cycle and cellular morphology of Sulfolobus at the single-cell level under the microscope. We would also love to catch a glimpse of Loki."
More information: Trends in Cell Biology, Dey et al.: "On the archaeal origins of eukaryotes" www.cell.com/trends/cell-biolo … 0962-8924(16)30002-2 , DOI: 10.1016/j.tcb.2016.03.009
Provided by Cell Press