What is CRISPR? And how can it be used to turn genes 'off'?

CRISPR systems have been a hot research topic since they were shown to have utility as genetic engineering tools in 2012. And they're often explained in a way that most folks can understand. But those explanations often overlook key details – like the fact that scientists are still in the process of discovering the fundamental rules of how these systems work.
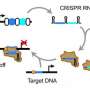
For example, here's a simplified explanation: CRISPR-Cas systems protect bacteria from invaders such as viruses. They do this by creating small strands of RNA that match DNA sequences specific to a given invader. When those CRISPR RNAs find a match, they unleash Cas proteins that chop up the invader's DNA, preventing it from replicating.
But of course it's more complex than that. For example, there are six different types of CRISPR systems (that we know of). One of the most widely-studied CRISPR systems is CRISPR-Cas9, which is a Type II CRISPR system.
But the most common CRISPR systems in nature are Type I. And new research from NC State is shedding light on some of the fundamental rules that govern Type I CRISPR systems – such as how long that CRISPR RNA can be, and how changing the length of the CRISPR RNA affects the behavior of the system.
To learn more, we talked to Chase Beisel and Michelle Luo, who recently published a paper on the work in Nucleic Acids Research, in collaboration with two groups at Montana State University. Beisel is an assistant professor of chemical and biomolecular engineering at NC State; Luo is a Ph.D. student in Beisel's lab.
The Abstract: Why are Type I CRISPR systems of particular interest?
Michelle Luo: As you mentioned, Type I systems are the most common type of CRISPR-Cas systems. They account for over half of known systems. This is of particular interest as we look into co-opting an organism's own system for other purposes. While CRISPR-Cas9 is undeniably a revolutionary genetic tool, it relies on importing this foreign Cas9 protein into an organism. This is a non-trivial task. However, if you use an organism's own CRISPR-Cas proteins, as shown in our earlier work, you can avoid the challenges of expressing a non-natural protein. Because Type I systems are so prevalent, they offer a promising route to explore how a natural CRISPR-Cas system can be exploited for other means.
The Abstract: In your recent work, you were evaluating how and whether you could modify the RNA in Type I CRISPR systems. Specifically, you were looking at whether you could modify the length of RNA in Type I CRISPR systems. Why would you want to change the length of the RNA?
Luo: Two years ago, a number of papers were published detailing the crystal structures of Type I protein complexes that bind and help degrade target DNA. These publications hinted at the CRISPR RNA serving as a scaffold to assemble the different proteins in the complex. In other words, the RNA serves as a framework for these proteins to grab onto. Thus, we hypothesized that if we changed the length of the CRISPR RNA, we could change the size and composition of the Type I protein complex, and possibly the complex's behavior.
The Abstract: How, or why, might expanding the protein complexes used in DNA recognition be useful?
Chase Beisel: Going into the project, we didn't know if the longer RNAs would allow the complex to even assemble, let alone function properly. We were surprised to find that the longer RNAs still formed a stable complex that could bind and direct the cutting of DNA. Because this complex is larger and recognizes a longer target sequence, we originally envisioned that the complex could be used for more specific DNA editing or for controlling gene expression.
The Abstract: When I think of CRISPR, I think of a system that either leaves DNA alone or cuts it up. What do you mean when you say that changing the length of the RNA is more effective at gene repression?
Luo: Your summary is on point. Normally, CRISPR-Cas systems survey the DNA landscape, and if they detects a target, they will cut up the DNA with tiny molecular scissors. If the target is not identified, the DNA will be left alone. Our earlier work demonstrated that we can prevent the cutting of the DNA by removing the scissors from the equation. We do this by deleting the cas3 gene from the genomic Type I locus. Now, instead of cutting the DNA, the CRISPR-Cas system simply binds the DNA. If we direct these modified systems to a gene, it will block the expression of that gene. Our most recent work shows that changing the length of the RNA can affect how strongly that silencing occurs. For certain regions, the longer the CRISPR RNA, the stronger the repression.
The Abstract: Does that make the CRISPR system more specific? I.e., does it allow the system to be more targeted in terms of the DNA it "attacks"?
Beisel: We wondered the same thing. We did in fact explore how longer RNAs impact specificity as part of the publication, although the results were mixed. On one hand, more of the RNA was involved in base pairing, where more base pairing would necessarily mean greater specificity. On the other hand, we found the longer RNAs were accommodating to mismatches with the target sequence, suggesting weaker specificity. In the end, more experiments will be needed to explore the question of specificity and how it impacts any downstream uses of Type I systems.
The Abstract: How might that gene repression function be used? Are there any potential applications?
Luo: Absolutely! This is particularly promising for metabolic engineering. If you want to make a microbial factory to produce a valuable product of interest, such as a biofuel, you have to alter the metabolism of an organism. This requires overexpressing genes that lead to production and turning off genes that compete with production. Our system allows researchers to turn off genes in a way that is potent, site-specific, reversible, and multiplexed. Our latest discovery suggests that you can fine-tune the extent of CRISPR-based gene repression simply by altering the length of the CRISPR RNA. That's what our recent paper in Nucleic Acids Research is about.
The Abstract: What are the future directions for this research?
Beisel: Aside from the applications Michelle mentioned, we're interested in why nature only uses RNA of a fixed length, given that longer RNAs make perfectly functional complexes. We're also interested in whether this phenomenon applies across the many different flavors of Type I systems, from those that use far fewer proteins in the complex to those found in organisms living at extreme temperatures.
More information: Kira S. Makarova et al. An updated evolutionary classification of CRISPR–Cas systems, Nature Reviews Microbiology (2015). DOI: 10.1038/nrmicro3569
Michelle L. Luo et al. The CRISPR RNA-guided surveillance complex inaccommodates extended RNA spacers, Nucleic Acids Research (2016). DOI: 10.1093/nar/gkw421
Journal information: Nature Reviews Microbiology , Nucleic Acids Research
Provided by North Carolina State University