Engineered E. coli vesicles act as vaccine deliverers
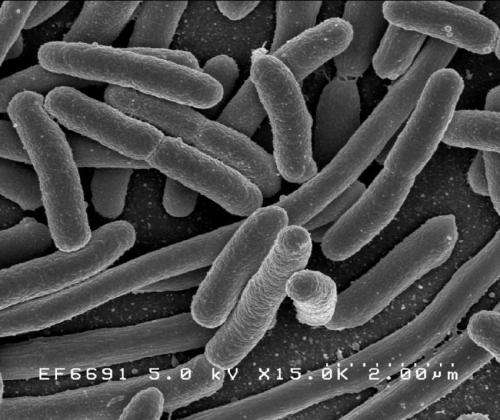
Mention E. coli and what pops into most people's heads are bacteria, tainted food, a rush to the hospital – basically, fear.
"E. coli gets a bad rap – and rightfully so," said Matthew DeLisa, the William L. Lewis Professor of Engineering.
But, as with many things, there are good varieties of E. coli and dangerous ones. And DeLisa, whose research focuses on developing tools for investigating and manipulating biological machinery directly in living bacterial cells, is working on a way to use a domesticated lab strain of E. coli to create and deliver vaccines.
A multi-institution effort – involving researchers from Cornell and the universities of Iowa, Texas and Georgia – has resulted in a paper that details how antigen-coated membrane vesicles derived from the surface of E. coli cells protected mice from a deadly pathogen, and how that system could work against other pathogens, as well.
The group's paper, "Outer membrane vesicles displaying engineered glycotypes elicit protective antibodies," was published online June 6 in the Proceedings of the National Academy of Sciences.
DeLisa, a member of the editorial board of the journal Cell Chemical Biology, has been engineering bacteria cells to do things they don't normally do for quite some time. His startup company, Glycobia, was the first to commercialize the use of engineered bacteria to make human glycoproteins – a protein modified with a carbohydrate attachment.
DeLisa and co-author David Putnam, associate professor in the Nancy E. and Peter C. Meinig School of Biomedical Engineering, have been working together for a decade on glycoproteins, or glycans – a protein with a carbohydrate attachment, which can be used to bind to certain protein receptor sites and, for example, block cancer cells from multiplying.
In this latest work, DeLisa's group has taken a similar approach to generating designer carbohydrate structures in E. coli, but instead of transferring the glycan to a protein, the cells assemble the glycan on a specific lipid carrier molecule. From there, the molecule is shuttled to the outer membrane of the E. coli cell, which then sheds small portions of that membrane.
"So you start with a 1-micron bacterial cell, and following vesiculation you get these small nanometer-scale spheres known as outer membrane vesicles (OMVs) that come off," DeLisa said. "Because they shed directly from the outer membrane of the cell, these OMVs are membrane-based nanostructures whose outer surfaces mimic the originating cell. So whatever is on the surface of the bacterial cell – say, an engineered glycolipid – becomes present on the surface of these vesicles."
These "conjugate" vaccines were injected into mice infected with the Francisella tularensis Schu S4 bacterium, the causative agent of tularemia and a feared bioweapon. That particular pathogen was chosen, DeLisa said, because of its potency and the lack of an existing vaccine.
"We might have chosen easier, lower-hanging fruit in terms of target pathogens," he said, "However, the lethality of Schu S4 – less than 10 colony-forming units are enough to kill you – made this organism a more interesting challenge."
Infected mice treated with the conjugate vesicle vaccines survived much longer than the control group. And in separate challenge experiments with a less virulent strain of Francisella, the vaccinated group saw 100 percent protection and survival of all mice.
A major advantage of OMVs as vaccine candidates is that they are potent adjuvants, which means they enhance the body's immune response to the co-delivered antigen.
"We think there's a real opportunity to further engineer these nanovesicles to go after other highly challenging vaccine targets," he said. "It's been fun and rewarding, because it brings together several research topics that we've been working on for a while now so we can leverage all that expertise."
More information: Linxiao Chen et al. Outer membrane vesicles displaying engineered glycotopes elicit protective antibodies, Proceedings of the National Academy of Sciences (2016). DOI: 10.1073/pnas.1518311113
Journal information: Cell Chemical Biology , Proceedings of the National Academy of Sciences
Provided by Cornell University