Researchers sequence first bedbug genome
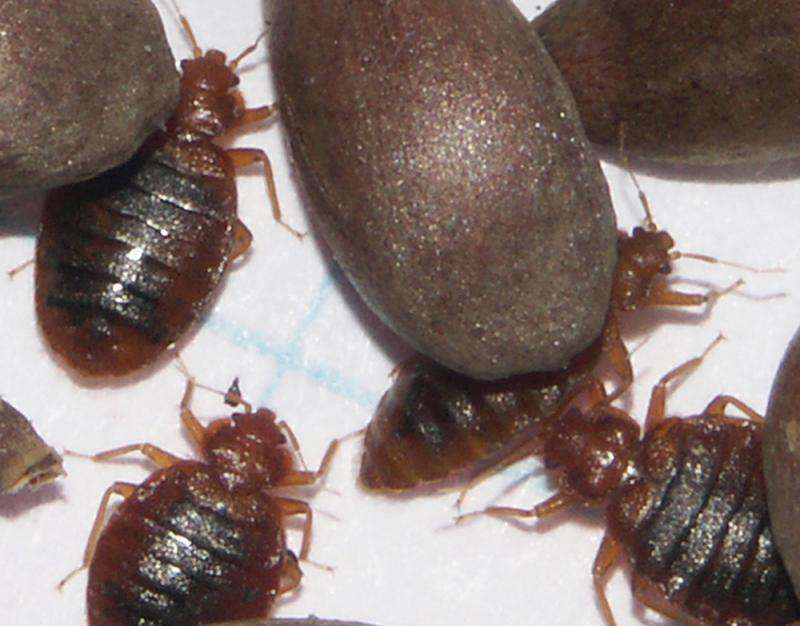

Scientists have assembled the first complete genome of one of humanity's oldest and least-loved companions: the bedbug. The new work, led by researchers at the American Museum of Natural History and Weill Cornell Medicine, and published Feb. 2 in Nature Communications, could help combat pesticide resistance in the unwelcome parasite. The data also provides a rich genetic resource for mapping bedbug activity in human hosts and in cities, including subways.
"Bedbugs are one of New York City's most iconic living fossils, along with cockroaches, meaning that their outward appearance has hardly changed throughout their long lineage," said corresponding author George Amato, director of the museum's Sackler Institute for Comparative Genomics. "But despite their static look, we know that they continue to evolve, mostly in ways that make it harder for humans to dissociate with them. This work gives us the genetic basis to explore the bedbug's basic biology and its adaptation to dense human environments."
The common bedbug (Cimex lectularius) has been coupled with humans for thousands of years. This species is found in temperate regions and prefers to feed on human blood. In recent decades, the prevalence of heated homes and global air travel has accelerated infestations in urban areas, where bedbugs have constant access to blood meals and opportunities to migrate to new hosts. A resurgence in bedbug infestations since the late 1990s is largely associated with the evolution of the insects' resistance to known pesticides, many of which are not suitable for indoor application.
"Bedbugs all but vanished from human lives in the 1940s because of the widespread use of DDT, but unfortunately, overuse contributed to resistance issues quite soon after that in bedbugs and other insect pests," said Louis Sorkin, an author on the paper and a senior scientific assistant in the museum's Division of Invertebrate Zoology. "Today, a very high percentage of bedbugs have genetic mutations that make them resistant to the insecticides that were commonly used to battle these urban pests. This makes the control of bedbugs extremely labor-intensive."
The researchers extracted DNA and RNA from preserved and living collections, including samples from a population that was first collected in 1973 and has been maintained by museum staff members since then. RNA was sampled from males and females representing each of the bug's six life stages, before and after blood meals, to paint a full picture of the bedbug genome.
"It's not enough to just sequence a genome, because by itself it does not tell the full story," said corresponding author Mark Siddall, a curator in the museum's Division of Invertebrate Zoology and Sackler Institute for Comparative Genomics. "In addition to the DNA, you want to get the RNA, or the expressed genes, and you want that not just from a single bedbug, but from both males and females at each part of the life cycle. Then you can really start asking questions about how certain genes relate to blood-feeding, insecticide resistance and other vital functions."
The researchers found that the number of genes was fairly consistent throughout the bedbug life cycle, but they observed notable changes in gene expression, especially after the first blood meal. Some genes, expressed only after the bedbug first drinks blood, are linked to insecticide resistance, including mechanisms that result in better detoxification and thicker chitin, or skin. This suggests that bedbugs are likely most vulnerable during the first nymph stage, potentially making it a good target for future insecticides. The scientists also identified three types of anticoagulant genes and their related proteins – characteristics consistent with a highly specialized blood feeder. When compared with 20 other arthropod genomes, the genome of the common bedbug shows close relationships to the kissing bug (Rhodnius prolixus), one of several vectors for Chagas disease, and the body louse (Pediculus humanus), which both have tight associations with humans.
The bedbug microbiome also contains more than 1,500 genes that map to more than 400 different species of bacteria, indicating that bedbugs harbor a rich suite of endosymbionts that are likely essential for their growth and reproduction. This indicates that antibiotics that attack bacteria beneficial to bed bugs (but that are non-essential to humans) could complement control of the insects via pesticides.
In addition, the study incorporated DNA data collected concurrently from more than 1,400 locations across New York City, including every subway station, to look at microbial diversity. For this work, the researchers used the new genomic data to focus exclusively on the diversity of bedbugs. They found differences in the genetic makeup of bedbugs that reside in different parts of the city, as measured by traces of DNA on east-west versus north-south subway lines, as well as between borough locations and among surfaces (e.g., benches vs. turnstiles). The findings suggest that areas of the city in close proximity to each other have bedbug populations that are related, and that bedbugs from one borough can be distinct from those in another borough. This kind of information can be used to map the pathways of migration of bedbug infestations in established and new urban environments.
"A great feature of metagenomics and microbiome data is its power to reveal new biology, since you can map previously unknown sequences to a new genome as soon as it is finished," said Christopher Mason, an associate professor of computational genomics in the Department of Physiology and Biophysics and the HRH Prince Alwaleed Bin Talal Bin Abdulaziz Al-Saud Institute for Computational Biomedicine at Weill Cornell Medicine and a senior author. "With every genome that is sequenced and annotated, the genetic understanding of the world around us becomes more in focus."
More information: Jeffrey A. Rosenfeld et al. Genome assembly and geospatial phylogenomics of the bed bug Cimex lectularius, Nature Communications (2016). DOI: 10.1038/ncomms10164
Journal information: Nature Communications
Provided by Cornell University